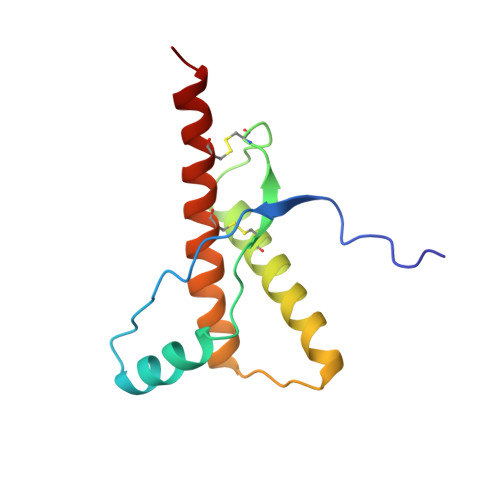
NMR Structure of a Variant Human Prion Protein with Two Disulfide Bridges
Zahn, R., Guntert, P., Von Schroetter, C., Wuthrich, K.(2003) J Mol Biology 326: 225
- PubMed: 12547204 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)01332-3
- Primary Citation Related Structures:
1H0L - PubMed Abstract:
The nuclear magnetic resonance structure of the globular domain with residues 121-230 of a variant human prion protein with two disulfide bonds, hPrP(M166C/E221C), shows the same global fold as wild-type hPrP(121-230). It contains three alpha-helices of residues 144-154, 173-194 and 200-228, an anti-parallel beta-sheet of residues 128-131 and 161-164, and the disulfides Cys166-Cys221 and Cys179-Cys214. The engineered extra disulfide bond in the presumed "protein X"-binding site is accommodated with slight, strictly localized conformational changes. High compatibility of hPrP with insertion of a second disulfide bridge in the protein X epitope was further substantiated by model calculations with additional variant structures. The ease with which the hPrP structure can accommodate a variety of locations for a second disulfide bond within the presumed protein X-binding epitope suggests a functional role for the extensive perturbation by a natural second disulfide bond of the corresponding region in the human doppel protein.
- Institut für Molekularbiologie und Biophysik, Eidgenössische Technische Hochschule Zürich, CH-8093, Zurich, Switzerland. rz@mol.biol.ethz.ch
Organizational Affiliation: