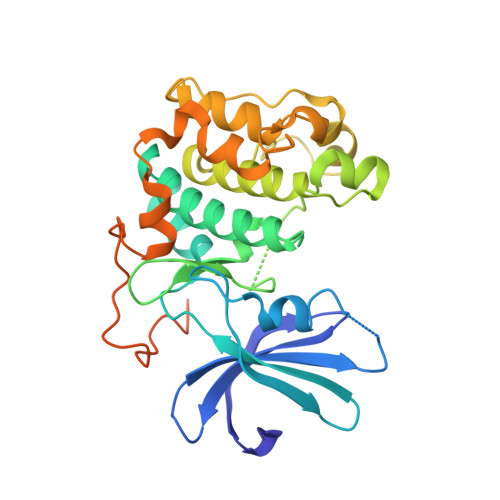
Molecular Mechanism for the Regulation of Protein Kinase B/Akt by Hydrophobic Motif Phosphorylation
Yang, J., Cron, P., Thompson, V., Good, V., Hess, D., Hemmings, B.A., Barford, D.(2002) Mol Cell 9: 1227
- PubMed: 12086620 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(02)00550-6
- Primary Citation Related Structures:
1GZK, 1GZN, 1GZO - PubMed Abstract:
Protein kinase B/Akt plays crucial roles in promoting cell survival and mediating insulin responses. The enzyme is stimulated by phosphorylation at two regulatory sites: Thr 309 of the activation segment and Ser 474 of the hydrophobic motif, a conserved feature of many AGC kinases. Analysis of the crystal structures of the unphosphorylated and Thr 309 phosphorylated states of the PKB kinase domain provides a molecular explanation for regulation by Ser 474 phosphorylation. Activation by Ser 474 phosphorylation occurs via a disorder to order transition of the alphaC helix with concomitant restructuring of the activation segment and reconfiguration of the kinase bilobal structure. These conformational changes are mediated by a phosphorylation-promoted interaction of the hydrophobic motif with a channel on the N-terminal lobe induced by the ordered alphaC helix and are mimicked by peptides corresponding to the hydrophobic motif of PKB and potently by the hydrophobic motif of PRK2.
- Section of Structural Biology, Institute of Cancer Research, Chester Beatty Laboratories, 237 Fulham Road, London SW3 6JB, UK.
Organizational Affiliation: