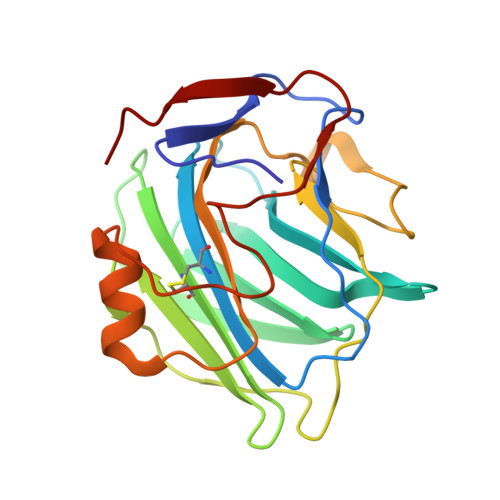
The Structures of Crystalline Complexes of Human Serum Amyloid P Component with its Carbohydrate Ligand, the Cyclic Pyruvate Acetal of Galactose
Thompson, D., Pepys, M.B., Tickle, I., Wood, S.P.(2002) J Mol Biology 320: 1081
- PubMed: 12126626 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00514-4
- Primary Citation Related Structures:
1GYK - PubMed Abstract:
Two monoclinic (P2(1)) crystal forms of human serum amyloid P component (SAP) in complex with the 4,6-pyruvate acetal of beta-D-galactose (MObetaDG) were prepared. Structure analysis by molecular replacement and refinement at 2.2A resolution revealed that crystal form 1 (a=95.76A, b=70.53A, c=103.41A, beta=96.80 degrees) contained a pentamer in the asymmetric unit with a structure very similar to that of the published search model. The mode of ligand co-ordination was also similar except that four of the five subunits showed bound ligand with an additional H-bond between O1 of the galactose and the side-chain of Lys79. One sub-unit showed no bound ligand and a vacant calcium site close to a crystal contact. The 2.6A resolution structure of crystal form 2 (a=118.60A, b=109.10A, c=120.80A and beta=95.16 degrees ) showed ten sub-units in the asymmetric unit, all with two bound calcium ions and ligand. The most extensive protein-protein interactions between pentamers describe an AB face-to-face interaction involving 15 ion pairs that sandwiches five molecules of bound MObetaDG at the interface.
- School of Biological Science, University of Southampton, Bassett Crescent East, Hampshire, UK. darren@soton.ac.uk
Organizational Affiliation: