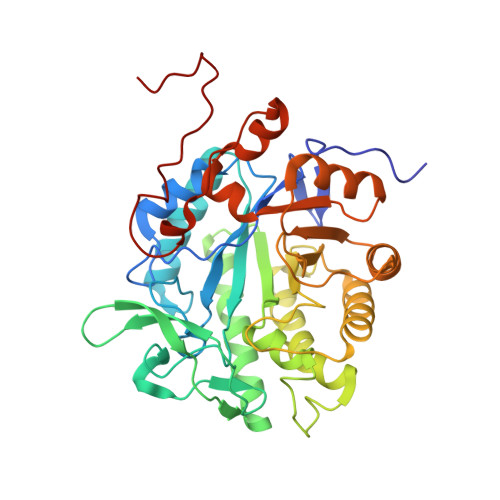
Crystal Structure of Bacterial Morphinone Reductase and Properties of the C191A Mutant Enzyme.
Barna, T.M., Messiha, H.L., Petosa, C., Bruce, N.C., Scrutton, N.S., Moody, P.C.E.(2002) J Biological Chem 277: 30976
- PubMed: 12048188
- DOI: https://doi.org/10.1074/jbc.M202846200
- Primary Citation Related Structures:
1GWJ - PubMed Abstract:
The crystal structure of the NADH-dependent bacterial flavoenzyme morphinone reductase (MR) has been determined at 2.2-A resolution in complex with the oxidizing substrate codeinone. The structure reveals a dimeric enzyme comprising two 8-fold beta/alpha barrel domains, each bound to FMN, and a subunit folding topology and mode of flavin-binding similar to that found in Old Yellow Enzyme (OYE) and pentaerythritol tetranitrate (PETN) reductase. The subunit interface of MR is formed by interactions from an N-terminal beta strand and helices 2 and 8 of the barrel domain and is different to that seen in OYE. The active site structures of MR, OYE, and PETN reductase are highly conserved reflecting the ability of these enzymes to catalyze "generic" reactions such as the reduction of 2-cyclohexenone. A region of polypeptide presumed to define the reducing coenzyme specificity is identified by comparison of the MR structure (NADH-dependent) with that of PETN reductase (NADPH-dependent). The active site acid identified in OYE (Tyr-196) and conserved in PETN reductase (Tyr-186) is replaced by Cys-191 in MR. Mutagenesis studies have established that Cys-191 does not act as a crucial acid in the mechanism of reduction of the olefinic bond found in 2-cyclohexenone and codeinone.
- Department of Biochemistry, University of Leicester, University Road, Leicester LE1 7RH, UK.
Organizational Affiliation: