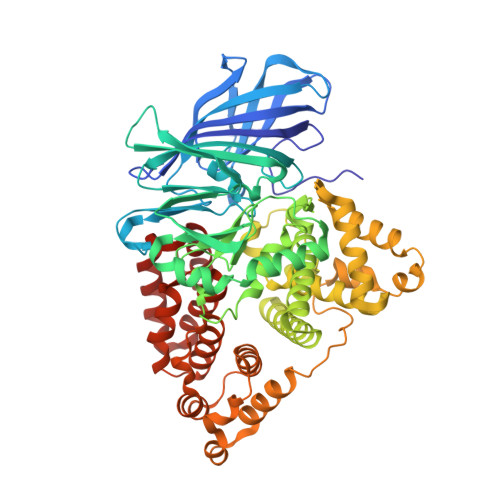
Leukotriene A4 Hydrolase: Selective Abrogation of Leukotriene B4 Formation by Mutation of Aspartic Acid 375
Rudberg, P.C., Tholander, F., Thunnissen, M.M.G.M., Samuelsson, B., Haeggstrom, J.Z.(2002) Proc Natl Acad Sci U S A 99: 4215
- PubMed: 11917124 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.072090099
- Primary Citation Related Structures:
1GW6 - PubMed Abstract:
Leukotriene A4 (LTA4, 5S-trans-5,6-oxido-7,9-trans-11,14-cis-eicosatetraenoic acid) hydrolase (LTA4H)/aminopeptidase is a bifunctional zinc metalloenzyme that catalyzes the final and rate-limiting step in the biosynthesis of leukotriene B4 (LTB4, 5S,12R-dihydroxy-6,14-cis-8,10-trans-eicosatetraenoic acid), a classical chemoattractant and immune modulating lipid mediator. Two chemical features are key to the bioactivity of LTB4, namely, the chirality of the 12R-hydroxyl group and the cis-trans-trans geometry of the conjugated triene structure. From the crystal structure of LTA4H, a hydrophilic patch composed of Gln-134, Tyr-267, and Asp-375 was identified in a narrow and otherwise hydrophobic pocket, believed to bind LTA4. In addition, Asp-375 belongs to peptide K21, a previously characterized 21-residue active site-peptide to which LTA4 binds during suicide inactivation. In the present report we used site-directed mutagenesis and x-ray crystallography to show that Asp-375, but none of the other candidate residues, is specifically required for the epoxide hydrolase activity of LTA4H. Thus, mutation of Asp-375 leads to a selective loss of the enzyme's ability to generate LTB4 whereas the aminopeptidase activity is preserved. We propose that Asp-375, possibly assisted by Gln-134, acts as a critical determinant for the stereoselective introduction of the 12R-hydroxyl group and thus the biological activity of LTB4.
- Department of Medical Biochemistry and Biophysics, Division of Chemistry II, Karolinska Institutet, S-171 77 Stockholm, Sweden.
Organizational Affiliation: