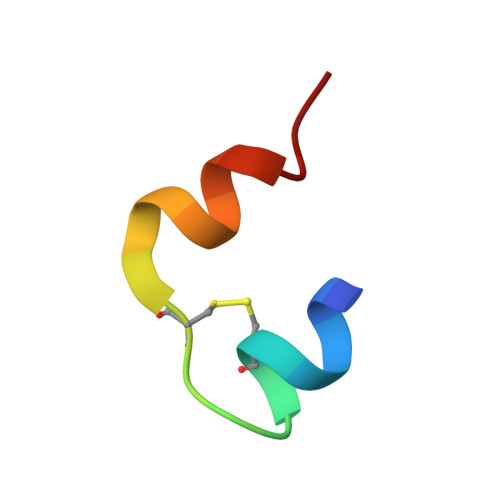
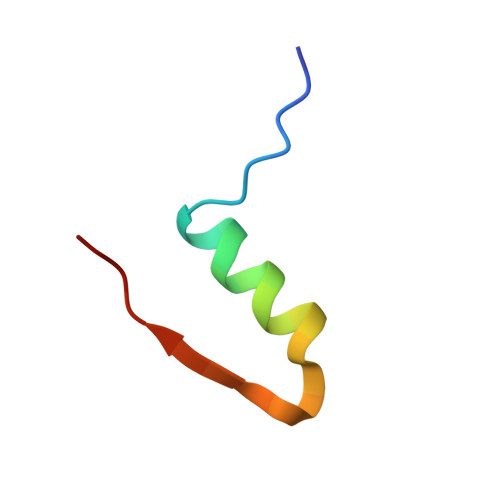
Insulin at Ph2: Structural Analysis of the Conditions Promoting Insulin Fibre Formation
Whittingham, J.L., Scott, D.J., Chance, K., Wilson, A., Finch, J., Brange, J., Dodson, G.G.(2002) J Mol Biology 318: 479
- PubMed: 12051853 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00021-9
- Primary Citation Related Structures:
1GUJ - PubMed Abstract:
When insulin solutions are subjected to acid, heat and agitation, the normal pattern of insulin assembly (dimers-->tetramers-->hexamers) is disrupted; the molecule undergoes conformational changes allowing it to follow an alternative aggregation pathway (via a monomeric species) leading to the formation of insoluble amyloid fibres. To investigate the effect of acid pH on the conformation and aggregation state of the protein, the crystal structure of human insulin at pH 2.1 has been determined to 1.6 A resolution. The structure reveals that the native fold is maintained at low pH, and that the molecule is still capable of forming dimers similar to those found in hexameric insulin structures at higher pH. Sulphate ions are incorporated into the molecule and the crystal lattice where they neutralise positive charges on the protein, stabilising its structure and facilitating crystallisation. The sulphate interactions are associated with local deformations in the protein, which may indicate that the structure is more plastic at low pH. Transmission electron microscopy analysis of insulin fibres reveals that the appearance of the fibres is greatly influenced by the type of acid employed. Sulphuric acid produces distinctive highly bunched, truncated fibres, suggesting that the sulphate ions have a sophisticated role to play in fibre formation, rather as they do in the crystal structure. Analytical ultracentrifugation studies show that in the absence of heating, insulin is predominantly dimeric in mineral acids, whereas in acetic acid the equilibrium is shifted towards the monomer. Hence, the effect of acid on the aggregation state of insulin is also complex. These results suggest that acid conditions increase the susceptibility of the molecule to conformational change and dissociation, and enhance the rate of fibrillation by providing a charged environment in which the attractive forces between the protein molecules is increased.
- York Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5DD, UK. jean@ysbl.york.ac.uk
Organizational Affiliation: