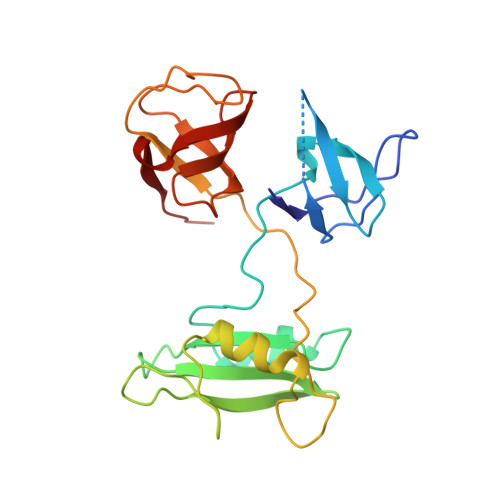
Crystal structure of the mammalian Grb2 adaptor.
Maignan, S., Guilloteau, J.P., Fromage, N., Arnoux, B., Becquart, J., Ducruix, A.(1995) Science 268: 291-293
- PubMed: 7716522 Search on PubMed
- DOI: https://doi.org/10.1126/science.7716522
- Primary Citation Related Structures:
1GRI - PubMed Abstract:
The mammalian growth factor receptor-binding protein Grb2 is an adaptor that mediates activation of guanine nucleotide exchange on Ras. Grb2 binds to the receptor through its SH2 domain and to the carboxyl-terminal domain of Son of sevenless through its two SH3 domains. It is thus a key element in the signal transduction pathway. The crystal structure of Grb2 was determined to 3.1 angstrom resolution. The asymmetric unit is composed of an embedded dimer. The interlaced junctions between the SH2 and SH3 domains bring the two adjacent faces of the SH3 domains in van der Waals contact but leave room for the binding of proline-rich peptides.
- Laboratoire de Biologie Structurale, Unite Mixte de Recherche CNRS-Université de Paris-Sud, Gif sur Yvette, France.
Organizational Affiliation: