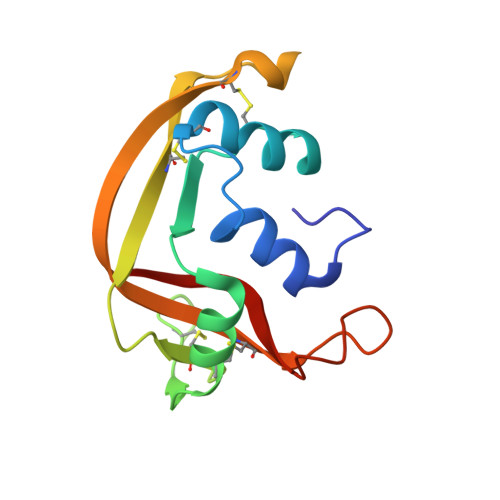
Atomic Resolution (0.98 A) Structure of Eosinophil-Derived Neurotoxin
Swaminathan, G.J., Holloway, D.E., Veluraja, K., Acharya, K.R.(2002) Biochemistry 41: 3341
- PubMed: 11876642 Search on PubMed
- DOI: https://doi.org/10.1021/bi015911f
- Primary Citation Related Structures:
1GQV - PubMed Abstract:
Human eosinophil-derived neurotoxin (EDN) is a small, basic protein that belongs to the ribonuclease A superfamily. EDN displays antiviral activity and causes the neurotoxic Gordon phenomenon when injected into rabbits. Although EDN and ribonuclease A have appreciable structural similarity and a conserved catalytic triad, their peripheral substrate-binding sites are not conserved. The crystal structure of recombinant EDN (rEDN) has been determined at 0.98 A resolution from data collected at a low temperature (100 K). We have refined the crystallographic model of the structure using anisotropic displacement parameters to a conventional R-factor of 0.116. This represents the highest resolution structure of rEDN determined to date and is only the second ribonuclease structure to be determined at a resolution greater than 1.0 A. The structure provides a detailed picture of the conformational freedom at the various subsites of rEDN, and the water structure accounts for more than 50% of the total solvent content of the unit cell. This information will be crucial for the design of tight-binding inhibitors to restrain the ribonucleolytic activity of rEDN.
- Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath BA2 7AY, U.K.
Organizational Affiliation: