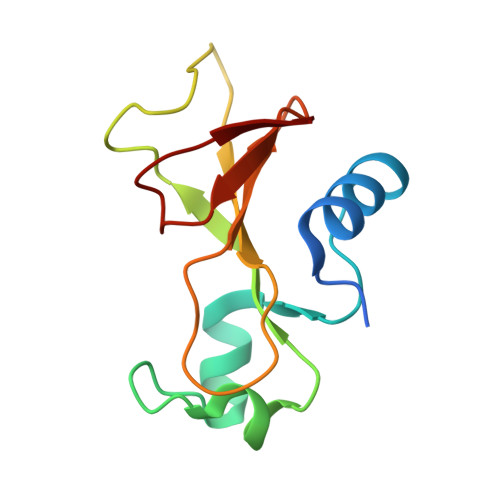
The Structure of Substrate-Free Microbial Ribonuclease Binase and of its Complexes with 3'Gmp and Sulfate Ions
Polyakov, K.M., Lebedev, A.A., Okorokov, A.L., Panov, K.I., Schulga, A.A., Pavlovsky, A.G., Karpeisky, M.Y.A., Dodson, G.G.(2002) Acta Crystallogr D Biol Crystallogr 58: 744
- PubMed: 11976484 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444902003207
- Primary Citation Related Structures:
1GOU, 1GOV, 1GOY - PubMed Abstract:
The structures of Bacillus intermedius ribonuclease (binase), an extracellular 109-residue enzyme, and its complexes with 3'GMP and sulfate ions were solved at 1.65 and 2.0 A, respectively. The structures were refined using REFMAC. The crystal of free binase belongs to the space group C2, whereas the crystals of complexes belong to the space group P2(1)2(1)2(1). In both crystal lattices the asymmetric unit contains two molecules which form an identical dimer. The structure of the dimer is such that only one of its subunits can bind the nucleotide in the 3'GMP-binase complex, where the guanyl base is located in the recognition loop of the enzyme. In both binase complex structures the phosphate group of 3'GMP or one of the sulfate ions make an electrostatic interaction with the binase molecule at the catalytic site. A second phosphate-binding site was found in the structures of the complexes at the cleft formed by the loop 34-39, the main chain of Arg82 and the side chain of Trp34. Comparison of the complex and unliganded enzyme crystal structures shows that there are some small but distinct differences in the specificity loop (56-62) and in the loops 34-39 and 99-104 associated with the binding of the nucleotide and sulfate ions.
- Institute of Crystallography, RAS, 107333 Moscow, Russia.
Organizational Affiliation: