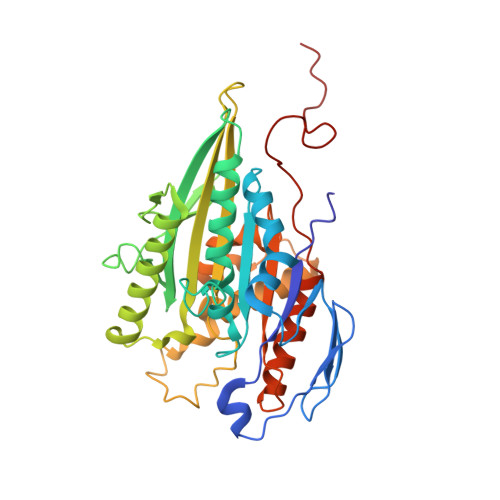
Structure of a Fast Kinesin: Implications for ATPase Mechanism and Interactions with Microtubules
Song, Y.-H., Marx, A., Muller, J., Woehlke, G., Schliwa, M., Krebs, A., Hoenger, A., Mandelkow, E.(2001) EMBO J 20: 6213
- PubMed: 11707393 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/20.22.6213
- Primary Citation Related Structures:
1GOJ - PubMed Abstract:
We determined the crystal structure of the motor domain of the fast fungal kinesin from Neurospora crassa (NcKin). The structure has several unique features. (i) Loop 11 in the switch 2 region is ordered and enables one to describe the complete nucleotide-binding pocket, including three inter-switch salt bridges between switch 1 and 2. (ii) Loop 9 in the switch 1 region bends outwards, making the nucleotide-binding pocket very wide. The displacement in switch 1 resembles that of the G-protein ras complexed with its guanosine nucleotide exchange factor. (iii) Loop 5 in the entrance to the nucleotide-binding pocket is remarkably long and interacts with the ribose of ATP. (iv) The linker and neck region is not well defined, indicating that it is mobile. (v) Image reconstructions of ice-embedded microtubules decorated with NcKin show that it interacts with several tubulin subunits, including a central beta-tubulin monomer and the two flanking alpha-tubulin monomers within the microtubule protofilament. Comparison of NcKin with other kinesins, myosin and G-proteins suggests that the rate-limiting step of ADP release is accelerated in the fungal kinesin and accounts for the unusually high velocity and ATPase activity.
- Max-Planck Unit for Structural Molecular Biology, D-22607 Hamburg, Germany. song@crystal.desy.de
Organizational Affiliation: