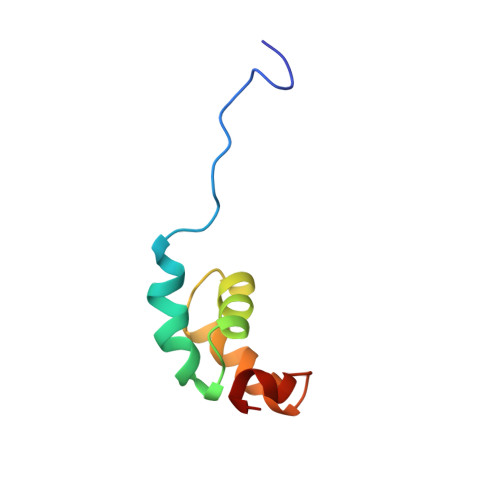
Structure of the C-Terminal Fg-Nucleoporin Binding Domain of Tap/Nxf1
Grant, R.P., Hurt, E., Neuhaus, D., Stewart, M.(2002) Nat Struct Biol 9: 247
- PubMed: 11875519 Search on PubMed
- DOI: https://doi.org/10.1038/nsb773
- Primary Citation Related Structures:
1GO5 - PubMed Abstract:
The vertebrate Tap protein is a member of the NXF family of shuttling transport receptors for nuclear export of mRNA. Tap has a modular structure, and its most C-terminal domain is important for binding to FG repeat-containing nuclear pore proteins (FG-nucleoporins) and is sufficient to mediate nuclear shuttling. We report the solution structure of this C-terminal domain, which is based on a distinctive arrangement of four alpha-helices and is joined to the next module by a flexible 12-residue Pro-rich linker. F617A Tap suppresses FG-nucleoporin binding by the most C-terminal domain that, together with the structure of the other modules from which Tap is constructed, provides a structural context for its nuclear shuttling function.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: