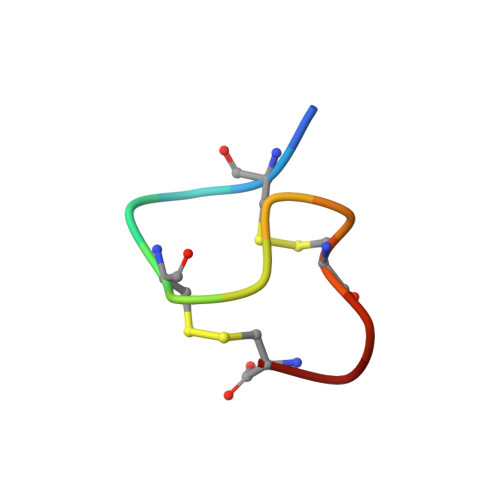
Determination of the solution structure of the peptide hormone guanylin: observation of a novel form of topological stereoisomerism.
Skelton, N.J., Garcia, K.C., Goeddel, D.V., Quan, C., Burnier, J.P.(1994) Biochemistry 33: 13581-13592
- PubMed: 7947768 Search on PubMed
- DOI: https://doi.org/10.1021/bi00250a010
- Primary Citation Related Structures:
1GNA, 1GNB - PubMed Abstract:
Guanylin is a 15 amino acid mammalian hormone containing two disulfide bonds. Guanylin shares sequence similarity with the bacterial heat-stable enterotoxin (STa) and is capable of binding to and stimulating the STa guanylyl cyclase receptor. Biologically active peptides have been prepared by two methods: (1) enzymatic treatment of a 99 residue proprotein (denoted proguanylin) expressed in Escherichia coli and (2) solid-phase chemical synthesis. Although both sources yield material that is pure by high-performance liquid chromatography and mass spectrometry, analysis by nuclear magnetic resonance (NMR) indicates that peptides from both sources contain two conformationally distinct species present in a 1:1 ratio. The chemical shift differences between the two species are large, allowing unambiguous sequential NMR assignments to be made for both sets of resonances. Exchange between the two forms was not observed even at 70 degrees C. Structural restraints have been generated from nuclear Overhauser effects and scalar coupling constants and used to calculate structures for both forms using distance geometry and restrained energy minimization. The resulting structures for the first isoform are well defined (root-mean-square deviation from the average structure for backbone atoms of 0.47 A) and adopt a right-handed spiral conformation, similar to that observed for heat stable enterotoxin. The second isoform is less well defined (root-mean-square deviation from the average structure for backbone atoms of 1.07 A) but clearly adopts a very different fold consisting of a left-hand spiral. The differences in structure suggest that the two forms may have very different affinities toward the STa receptor. The observation of such isomerism has important implications for the common practice of introducing multiple disulfide bonds into small peptides to limit conformational flexibility and enhance bioactivity.
- Department of Protein Engineering, Genentech, Inc., South San Francisco, California 94080.
Organizational Affiliation: