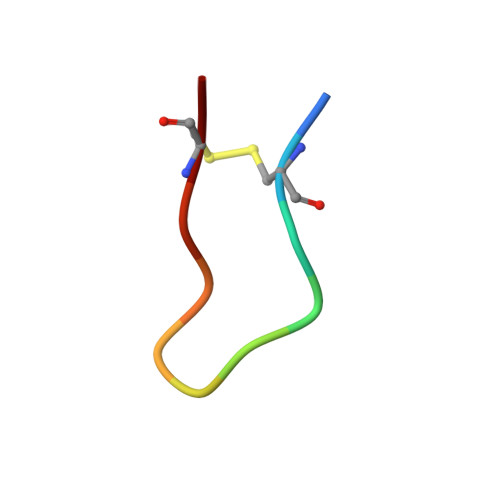
The (1)H-NMR solution structure of the antitryptic core peptide of Bowman-Birk inhibitor proteins: a minimal canonical loop.
Brauer, A.B., Kelly, G., Matthews, S.J., Leatherbarrow, R.J.(2002) J Biomol Struct Dyn 20: 59-70
- PubMed: 12144352 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.2002.10506822
- Primary Citation Related Structures:
1GM2 - PubMed Abstract:
Bowman-Birk inhibitor (BBI) proteins contain an inhibitory motif comprising a disulfide-bonded sequence that interacts with serine proteinases. Recently, a small 14-residue peptide from sunflowers (SFTI-1), which has potent anti-trypsin activity, has been found to have the same motif. However, this peptide also has an unusual head-to-tail cyclisation. To address the role of the core inhibitory sequence itself, we have solved the (1)H-NMR solution structure of an antitryptic 11-residue cyclic peptide that corresponds to the core reactive site loops of both SFTI-1 and Bowman-Birk inhibitor proteins. A comparison is made between the secondary chemical shifts found in this family and the canonical regions of several other inhibitors, giving some insight into relative flexibility and hydrogen bonding patterns in these inhibitors. The solution structure of the core peptide in isolation is found to retain essentially the same three-dimensional arrangement of both backbone and side chains as observed in larger antitryptic BBI and SFTI-1 fragments as well as in the complete proteins. The retention of the canonical conformation in the core peptide explains the peptids inhibitory potency. It therefore represents a minimization of both the BBI and SFTI-1 sequences. We conclude that the core peptide is a conformationally defined, canonical scaffold, which can serve as a minimal platform for the engineering of biological activity.
- Department of Chemistry, Imperial College of Science, Technology and Medicine, South Kensington, London, SW7 2AY, U.K.
Organizational Affiliation: