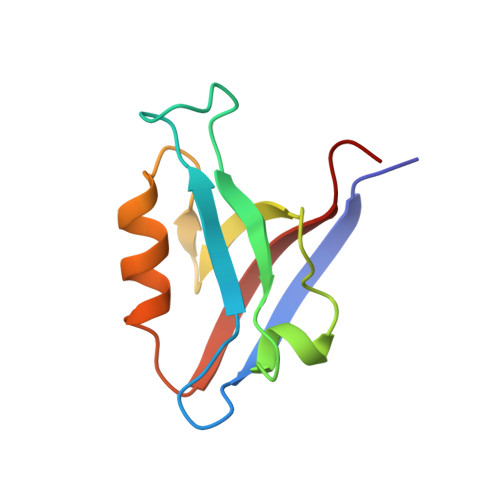
Structure, dynamics and binding characteristics of the second PDZ domain of PTP-BL.
Walma, T., Spronk, C.A., Tessari, M., Aelen, J., Schepens, J., Hendriks, W., Vuister, G.W.(2002) J Mol Biology 316: 1101-1110
- PubMed: 11884147 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2002.5402
- Primary Citation Related Structures:
1GM1 - PubMed Abstract:
The PDZ domains of the protein tyrosine phosphatase PTP-BL mediate interactions by binding to specific amino acid sequences in target proteins. The solution structure of the second PDZ domain of PTP-BL, PDZ2, displays a compact fold with six beta strands and two alpha-helices. A unique feature of this domain compared to the canonical PDZ fold is an extended flexible loop at the base of the binding pocket, termed L1, that folds back onto the protein backbone, a feature that is shared by both the murine and human orthologues. The structure of PDZ2 differs significantly from the orthologous human structure. A comparison of structural quality indicators clearly demonstrates that the PDZ2 ensemble is statistically more reasonable than that of the human orthologue. The analysis of (15)N relaxation data for PDZ2 shows a normal pattern, with more rigid secondary structures and more flexible loop structures. Close to the binding pocket, Leu85 and Thr88 display greater mobility when compared to surrounding residues. Peptide binding studies demonstrated a lack of interaction between murine PDZ2 and the C terminus of the murine Fas/CD95 receptor, suggesting that the Fas/CD95 receptor is not an in vivo target for PDZ2. In addition, PDZ2 specifically binds the C termini of both human Fas/CD95 receptor and the RIL protein, despite RIL containing a non-canonical PDZ-interacting sequence of E-x-V. A model of PDZ2 with the RIL peptide reveals that the PDZ2 binding pocket is able to accommodate the bulkier side-chain of glutamic acid while maintaining crucial protein to peptide hydrogen bond interactions.
- NSR-RIM Center, Department of Biophysical Chemistry, University of Nijmegen, Toernooiveld 1, 6525 ED Nijmegen, The Netherlands.
Organizational Affiliation: