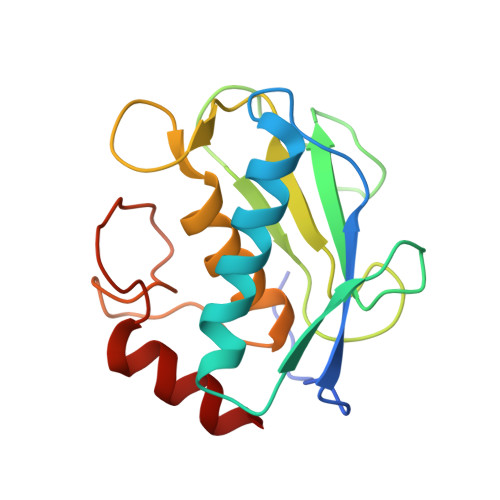
Crystal Structure of Mmp9 in Complex with a Reverse Hydroxamate Inhibitor
Rowsell, S., Hawtin, P., Minshull, C.A., Jepson, H., Brockbank, S., Barratt, D., Slater, A.M., Mcpheat, W., Waterson, D., Henney, A., Pauptit, R.A.(2002) J Mol Biology 319: 173
- PubMed: 12051944 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00262-0
- Primary Citation Related Structures:
1GKC, 1GKD - PubMed Abstract:
Matrix metalloproteinases (MMPs) and their inhibitors are important in connective tissue re-modelling in diseases of the cardiovascular system, such as atherosclerosis. Various members of the MMP family have been shown to be expressed in atherosclerotic lesions, but MMP9 is consistently seen in inflammatory atherosclerotic lesions. MMP9 over-expression is implicated in the vascular re-modelling events preceding plaque rupture (the most common cause of acute myocardial infarction). Reduced MMP9 activity, either by genetic manipulation or through pharmacological intervention, has an impact on ventricular re-modelling following infarction. MMP9 activity may therefore represent a key mechanism in the pathogenesis of heart failure. We have determined the crystal structure, at 2.3 A resolution, of the catalytic domain of human MMP9 bound to a peptidic reverse hydroxamate inhibitor as well as the complex of the same inhibitor bound to an active-site mutant (E402Q) at 2.1 A resolution. MMP9 adopts the typical MMP fold. The catalytic centre is composed of the active-site zinc ion, co-ordinated by three histidine residues (401, 405 and 411) and the essential glutamic acid residue (402). The main differences between the catalytic domains of various MMPs occur in the S1' subsite or selectivity pocket. The S1' specificity site in MMP9 is perhaps best described as a tunnel leading toward solvent, as in MMP2 and MMP13, as opposed to the smaller pocket found in fibroblast collagenase and matrilysin. The present structure enables us to aid the design of potent and specific inhibitors for this important cardiovascular disease target.
- AstraZeneca, Mereside, Alderley Park, Macclesfield, Cheshire SK10 4TG, UK.
Organizational Affiliation: