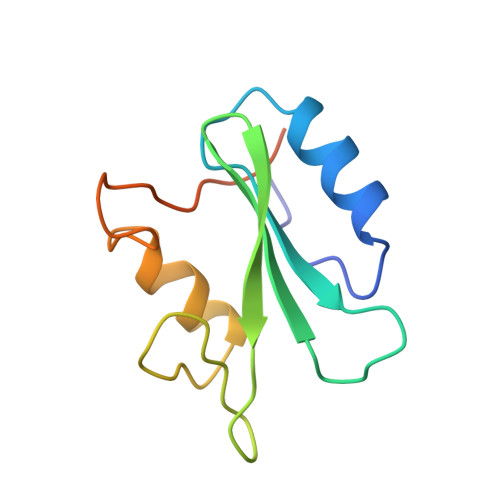
Nuclear magnetic resonance solution structure of the growth factor receptor-bound protein 2 Src homology 2 domain.
Thornton, K.H., Mueller, W.T., McConnell, P., Zhu, G., Saltiel, A.R., Thanabal, V.(1996) Biochemistry 35: 11852-11864
- PubMed: 8794768 Search on PubMed
- DOI: https://doi.org/10.1021/bi952615s
- Primary Citation Related Structures:
1GHU - PubMed Abstract:
A family of NMR solution structures of the growth factor receptor-bound protein 2 (Grb2) SH2 domain has been determined by heteronuclear multidimensional NMR. Proton, nitrogen, and carbon chemical shift assignments have been made for the SH2 domain of Grb2. Assignments were made from a combination of homonuclear two-dimensional and 15N- and 13C-edited three-dimensional spectra at pH 6.2 and 298 K. Structure-induced proton and carbon secondary shifts were calculated and used to facilitate the spectral assignment process. NOE, scalar coupling, secondary chemical shift, and amide proton exchange data were used to characterize the secondary structural elements and hydrogen-bonding network in the Grb2 SH2 domain. The three-dimensional structure of the Grb2 SH2 domain was calculated using 1112 restraints obtained from NOE, coupling constant, and amide proton exchange data. The rmsd for the 24 calculated structures to the mean structure of the Grb2 SH2 domain was 0.75 A for backbone and 1.28 A for all heavy atoms. The three-dimensional fold of the Grb2 SH2 domain is similar to that observed for other SH2 domains and consists of two alpha-helical segments and eight beta-strands, six strands that make up two contiguous antiparallel beta-sheets, and two strands that form two short parallel beta-sheets. The structure of the phosphotyrosine binding pocket of Grb2 is similar to that observed for other SH2 domains. The hydrophobic binding pocket of Grb2 is similar to that observed for Src with the exception that tryptophan 121 of Grb2 occupies part of the pY+3 binding pocket. Structural implications for the Grb2 SH2 domain selectivity at the pY+2 and pY+3 sites are discussed.
- Department of Chemistry, Parke-Davis Pharmaceutical Research, Division of Warner-Lambert Company, Ann Arbor, Michigan, USA.
Organizational Affiliation: