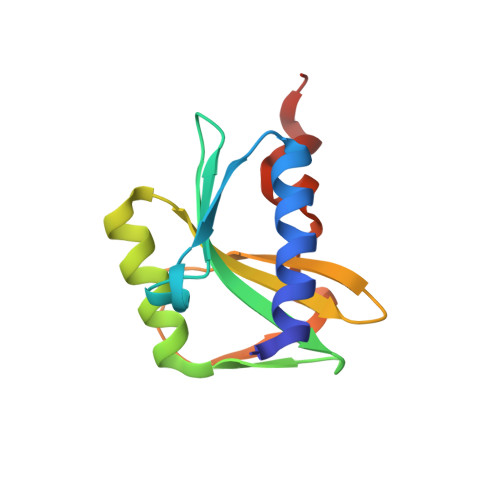
Crystal structure of the archaeal holliday junction resolvase Hjc and implications for DNA recognition.
Nishino, T., Komori, K., Tsuchiya, D., Ishino, Y., Morikawa, K.(2001) Structure 9: 197-204
- PubMed: 11286886 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00576-7
- Primary Citation Related Structures:
1GEF - PubMed Abstract:
Homologous recombination is a crucial mechanism in determining genetic diversity and repairing damaged chromosomes. Holliday junction is the universal DNA intermediate whose interaction with proteins is one of the major events in the recombinational process. Hjc is an archaeal endonuclease, which specifically resolves the junction DNA to produce two separate recombinant DNA duplexes. The atomic structure of Hjc should clarify the mechanisms of the specific recognition with Holliday junction and the catalytic reaction. The crystal structure of Hjc from the hyperthermophilic archaeon Pyrococcus furiosus has been determined at 2.0 A resolution. The active Hjc molecule forms a homodimer, where an extensive hydrophobic interface tightly assembles two subunits of a single compact domain. The folding of the Hjc subunit is clearly different from any other Holliday junction resolvases thus far known. Instead, it resembles those of type II restriction endonucleases, including the configurations of the active site residues, which constitute the canonical catalytic motifs. The dimeric Hjc molecule displays an extensive basic surface on one side, which contains many conserved amino acids, including those in the active site. The architectural similarity of Hjc to restriction endonucleases allowed us to construct a putative model of the complex with Holliday junction. This model accounts for how Hjc recognizes and resolves the junction DNA in a specific manner. Mutational and biochemical analyses highlight the importance of some loops and the amino terminal region in interaction with DNA.
- Department of Structural Biology, Biomolecular Engineering Research Institute, 6-2-3 Furuedai, Suita, Osaka 565-0874, Japan.
Organizational Affiliation: