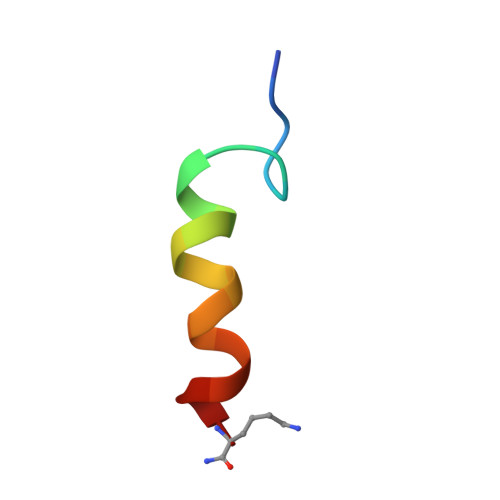
Conformation of a peptide ligand bound to its G-protein coupled receptor.
Inooka, H., Ohtaki, T., Kitahara, O., Ikegami, T., Endo, S., Kitada, C., Ogi, K., Onda, H., Fujino, M., Shirakawa, M.(2001) Nat Struct Biol 8: 161-165
- PubMed: 11175907 Search on PubMed
- DOI: https://doi.org/10.1038/84159
- Primary Citation Related Structures:
1GEA - PubMed Abstract:
Many peptide hormones elicit a wide array of physiological effects by binding to G-protein coupled receptors. We have determined the conformation of pituitary adenylate cyclase activating polypeptide, PACAP(1--21)NH(2), bound to a PACAP-specific receptor by NMR spectroscopy. Residues 3--7 form a unique beta-coil structure that is preceded by an N-terminal extended tail. This beta-coil creates a patch of hydrophobic residues that is important for receptor binding. In contrast, the C-terminal region (residues 8--21) forms an alpha-helix, similar to that in the micelle-bound PACAP. Thus, the conformational difference between PACAP in the receptor-bound and the micelle-bound states is limited to the N-terminal seven residues. This observation is consistent with the two-step ligand transportation model in which PACAP first binds to the membrane nonspecifically and then diffuses two-dimensionally in search of its receptor; a conformational change at the N-terminal region then allows specific interactions between the ligand and the receptor.
- Graduate School of Biological Sciences, Nara Institute of Science and Technology, 8916-5 Takayama, Ikoma, Nara 630-0101, Japan. Inooka_Hiroshi@takeda.co.jp
Organizational Affiliation: