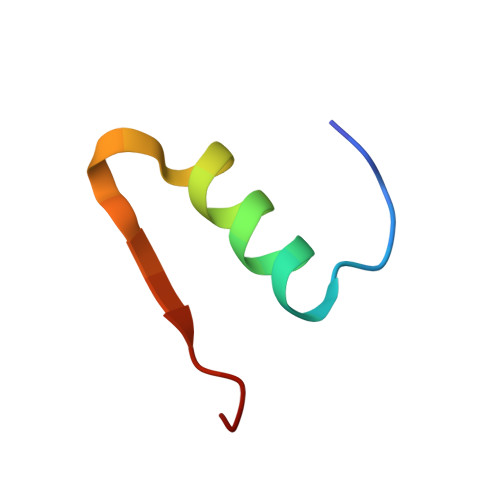
Phase changes in T(3)R(3)(f) human insulin: temperature or pressure induced?
Smith, G.D., Pangborn, W.A., Blessing, R.H.(2001) Acta Crystallogr D Biol Crystallogr 57: 1091-1100
- PubMed: 11468392 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444901007685
- Primary Citation Related Structures:
1G7A, 1G7B - PubMed Abstract:
The structure of T(3)R(3) hexameric human insulin has been determined at 100 K from two different crystals at 1.2 and 1.3 A resolution and refined to residuals of 0.169 and 0.176, respectively. Owing to a phase change, the c axis is double its room-temperature value and the asymmetric unit contains two independent TR(f) insulin dimers. Compared with the orientation in the room-temperature structure, one dimer undergoes a rotation about the c axis of -5 degrees, while the second is rotated +4 degrees. A superposition of the backbone atoms of the two independent dimers shows that the C(alpha) atoms of five residues within the R(f)-state monomers are displaced by more than 1.0 A; smaller displacements are observed for the T-state monomers. Four zinc ions lie on the crystallographic threefold axis and each forms bonds to three symmetry-related HisB10 N(varepsilon2) atoms from the T- and R(f)-state trimers. While three of the zinc ions are tetrahedrally coordinated with a chloride ion completing the coordination sphere, mixed tetrahedral/octahedral coordination is observed for one of the T-state zinc ions. The three symmetry-related "phenolic binding sites" in one hexamer contain water molecules and a glycerol molecule, but the same sites in the second hexamer are occupied by a zinc ion coordinated to an alternate conformation of HisB10, a symmetry-related HisB5 and two chloride ions. Two additional and partially occupied zinc ion sites are observed at the interface between the two independent dimers. One zinc ion is coordinated by a T-state HisB5 of one dimer, an R-state HisB5 of the second dimer and two water molecules; the second zinc ion is coordinated by an alternate side-chain conformation of the T-state HisB5 and three water molecules. The carboxyl group of one GluB13 side chain, which exists in two discrete conformations, appears to be protonated, because short contacts exist to a second carboxyl group or to a carbonyl O atom.
- Hauptman-Woodward Medical Research Institute, 73 High Street, Buffalo, NY 14203, USA. smith@hwi.buffalo.edu
Organizational Affiliation: