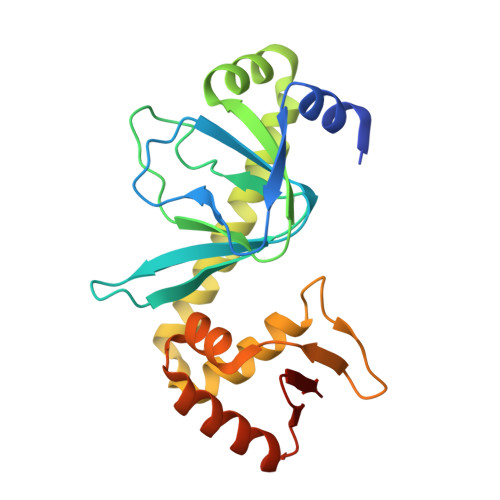
Modeling the cAMP-induced allosteric transition using the crystal structure of CAP-cAMP at 2.1 A resolution.
Passner, J.M., Schultz, S.C., Steitz, T.A.(2000) J Mol Biology 304: 847-859
- PubMed: 11124031 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4231
- Primary Citation Related Structures:
1G6N - PubMed Abstract:
After an allosteric transition produced by the binding of cyclic AMP (cAMP), the Escherichia coli catabolite gene activator protein (CAP) binds DNA specifically and activates transcription. The three-dimensional crystal structure of the CAP-cAMP complex has been refined at 2.1 A resolution, thus enabling a better evaluation of the structural basis for CAP phenotypes, the interactions of cAMP with CAP and the roles played by water structure. A review of mutational analysis of CAP together with the additional structural information presented here suggests a possible mechanism for the cAMP-induced allostery required for DNA binding and transcriptional activation. We hypothesize that cAMP binding may reorient the coiled-coil C-helices, which provide most of the dimer interface, thereby altering the relative positions of the DNA-binding domains of the CAP dimer. Additionally, cAMP binding may cause a further rearrangement of the DNA-binding and cAMP-binding domains of CAP via a flap consisting of beta-strands 4 and 5 which lies over the cAMP.
- Department of Molecular Biophysics and Biochemistry, Mount Sinai Schoolof Medicine, New York, NY 10029, USA. Passner@inka.mssm.edu
Organizational Affiliation: