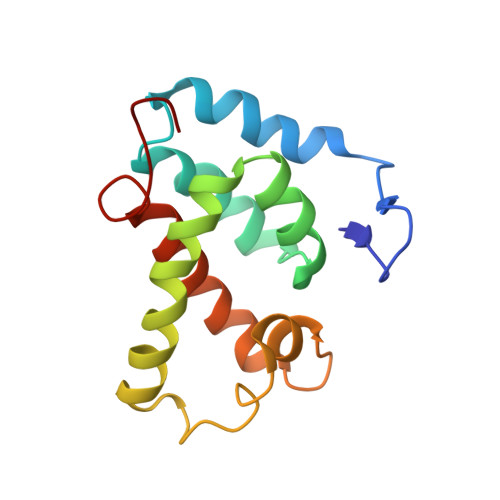
Structural analysis of the N-terminal domain of the human T-cell leukemia virus capsid protein.
Cornilescu, C.C., Bouamr, F., Yao, X., Carter, C., Tjandra, N.(2001) J Mol Biology 306: 783-797
- PubMed: 11243788 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4395
- Primary Citation Related Structures:
1G03 - PubMed Abstract:
The N-terminal domain of the retroviral capsid (CA) protein is one of the least conserved regions encoded in the genome. Surprisingly, the three-dimensional structures of the CA from different genera exhibit alpha-helical structural features that are highly conserved. The N-terminal residues of the human immunodeficiency virus type 1 (HIV-1) and Rous sarcoma virus (RSV) capsid proteins form a beta-hairpin. To determine if this feature is conserved in the retroviral family, we cloned, expressed, purified, and solved the structure of a N-terminal 134 amino acid fragment (CA(134)) from the human T-cell leukemia virus type 1 (HTLV-I) using high resolution nuclear magnetic resonance (NMR) spectroscopy. The CA(134) fragment contains an N-terminal beta-hairpin and a central coiled-coil-like structure composed of six alpha-helices. The N-terminal Pro1 residue contacts Asp54 in the helical cluster through a salt bridge. Thus, the beta-hairpin is conserved and the helical cluster is structurally similar to other retroviral CA domains. However, although the same Asp residue defines the orientation of the hairpin in both the HTLV-1 and HIV-1 CA proteins, the HTLV-I hairpin is oriented away, rather than towards, the helical core. Significant differences were also detected in the spatial orientation and helical content of the long centrally located loop connecting the helices in the core. It has been proposed that the salt bridge allows the formation of a CA-CA interface that is important for the assembly of the conical cores that are characteristic of HIV-1. As HTLV-I forms spherical cores, the salt-bridge feature is apparently not conserved for this function although its role in determining the orientation of the beta-hairpin may be critical, along with the central loop. Comparison of three-dimensional structures is expected to elucidate the relationships between the retroviral capsid protein structure and its function.
- Laboratory of Biophysical Chemistry, Bldg. 3, NHLBI, NIH, Bethesda, MD, 20892-0380, USA
Organizational Affiliation: