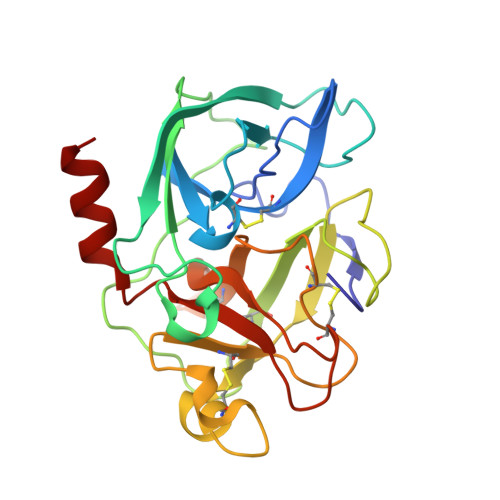
The crystal structure of the complex of non-peptidic inhibitor of human neutrophil elastase ONO-6818 and porcine pancreatic elastase.
Odagaki, Y., Ohmoto, K., Matsuoka, S., Hamanaka, N., Nakai, H., Toda, M., Katsuya, Y.(2001) Bioorg Med Chem 9: 647-651
- PubMed: 11310599 Search on PubMed
- DOI: https://doi.org/10.1016/s0968-0896(00)00277-7
- Primary Citation Related Structures:
1FZZ - PubMed Abstract:
The crystal structure of a new inhibitor of human neutrophil elastase (HNE), N-[2-[5-(tert-butyl)-1,3,4-oxadiazol-2-yl]-(IRS)-1-(methylethyl)-2-oxoethyl]-2-(5-amino-6-oxo-2-phenyl-6H-pyrimidin-1-ly)acetamide (ONO-6818, 1) complexed to porcine pancreatic elastase (PPE) has been determined at 1.86 A resolution. Analytical results provided evidence of a 1:1 complex in which the electrophilic ketone of 1 covalently bound to O gamma of Ser195 at the active site of PPE. The role of the unique electron-withdrawing ketone of 1 has been elucidated.
- Minase Research Institute, Ono Pharmaceutical Co., Ltd. Mishima-Gun, Osaka, Japan. odagaki@ono.co.jp
Organizational Affiliation: