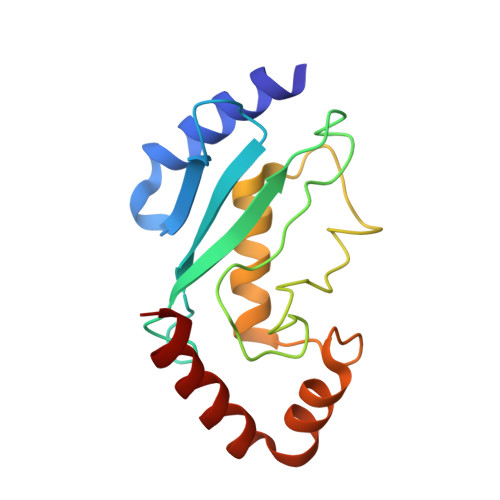
Structure of a conjugating enzyme-ubiquitin thiolester intermediate reveals a novel role for the ubiquitin tail.
Hamilton, K.S., Ellison, M.J., Barber, K.R., Williams, R.S., Huzil, J.T., McKenna, S., Ptak, C., Glover, M., Shaw, G.S.(2001) Structure 9: 897-904
- PubMed: 11591345 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00657-8
- Primary Citation Related Structures:
1FXT, 1FZY - PubMed Abstract:
Ubiquitin-conjugating enzymes (E2s) are central enzymes involved in ubiquitin-mediated protein degradation. During this process, ubiquitin (Ub) and the E2 protein form an unstable E2-Ub thiolester intermediate prior to the transfer of ubiquitin to an E3-ligase protein and the labeling of a substrate for degradation. A series of complex interactions occur among the target substrate, ubiquitin, E2, and E3 in order to efficiently facilitate the transfer of the ubiquitin molecule. However, due to the inherent instability of the E2-Ub thiolester, the structural details of this complex intermediate are not known. A three-dimensional model of the E2-Ub thiolester intermediate has been determined for the catalytic domain of the E2 protein Ubc1 (Ubc1(Delta450)) and ubiquitin from S. cerevisiae. The interface of the E2-Ub intermediate was determined by kinetically monitoring thiolester formation by 1H-(15)N HSQC spectra by using combinations of 15N-labeled and unlabeled Ubc1(Delta450) and Ub proteins. By using the surface interface as a guide and the X-ray structures of Ub and the 1.9 A structure of Ubc1(Delta450) determined here, docking simulations followed by energy minimization were used to produce the first model of a E2-Ub thiolester intermediate. Complementary surfaces were found on the E2 and Ub proteins whereby the C terminus of Ub wraps around the E2 protein terminating in the thiolester between C88 (Ubc1(Delta450)) and G76 (Ub). The model supports in vivo and in vitro experiments of E2 derivatives carrying surface residue substitutions. Furthermore, the model provides insights into the arrangement of Ub, E2, and E3 within a ternary targeting complex.
- Department of Biochemistry, The University of Alberta, Edmonton, Alberta T6G 2H7, Canada.
Organizational Affiliation: