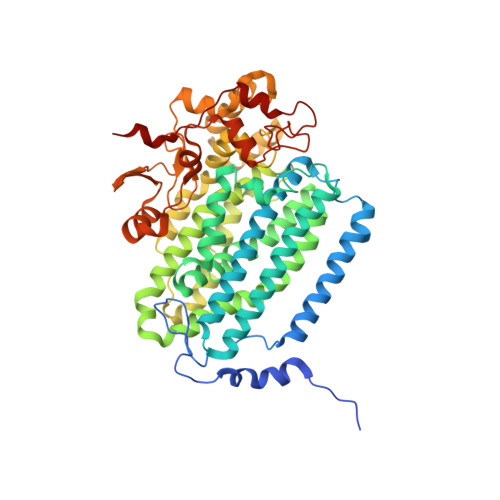
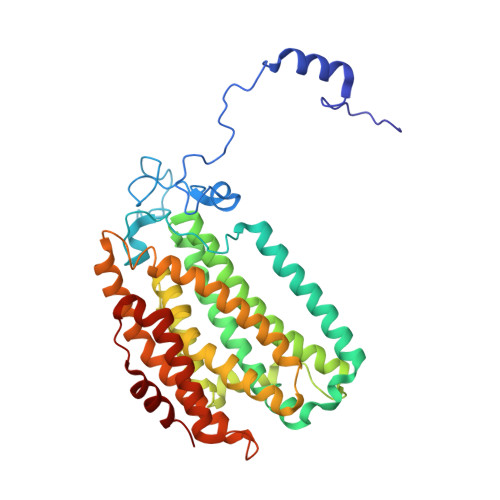
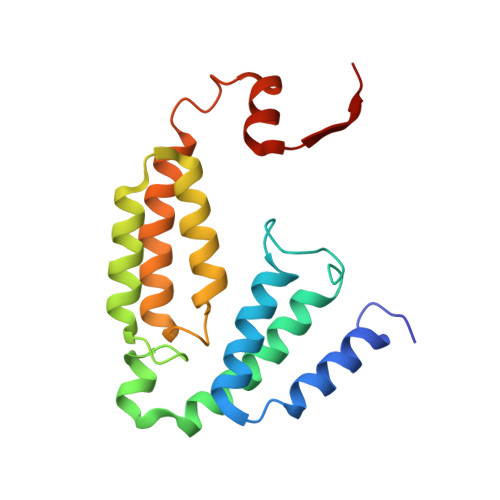
X-ray crystal structure of alcohol products bound at the active site of soluble methane monooxygenase hydroxylase.
Whittington, D.A., Sazinsky, M.H., Lippard, S.J.(2001) J Am Chem Soc 123: 1794-1795
- PubMed: 11456795 Search on PubMed
- DOI: https://doi.org/10.1021/ja0031725
- Primary Citation Related Structures:
1FZ6, 1FZ7 - Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: