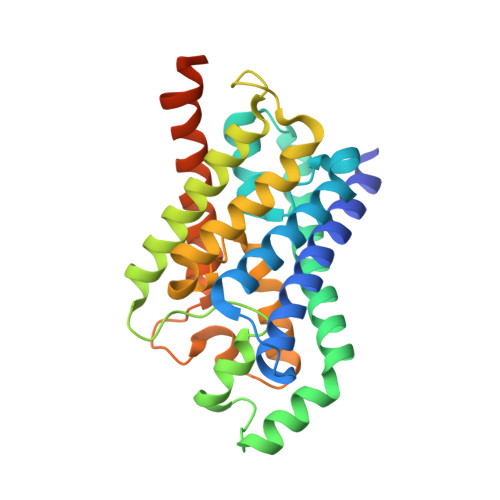
Structure of a glycerol-conducting channel and the basis for its selectivity.
Fu, D., Libson, A., Miercke, L.J., Weitzman, C., Nollert, P., Krucinski, J., Stroud, R.M.(2000) Science 290: 481-486
- PubMed: 11039922 Search on PubMed
- DOI: https://doi.org/10.1126/science.290.5491.481
- Primary Citation Related Structures:
1FX8 - PubMed Abstract:
Membrane channel proteins of the aquaporin family are highly selective for permeation of specific small molecules, with absolute exclusion of ions and charged solutes and without dissipation of the electrochemical potential across the cell membrane. We report the crystal structure of the Escherichia coli glycerol facilitator (GlpF) with its primary permeant substrate glycerol at 2.2 angstrom resolution. Glycerol molecules line up in an amphipathic channel in single file. In the narrow selectivity filter of the channel the glycerol alkyl backbone is wedged against a hydrophobic corner, and successive hydroxyl groups form hydrogen bonds with a pair of acceptor, and donor atoms. Two conserved aspartic acid-proline-alanine motifs form a key interface between two gene-duplicated segments that each encode three-and-one-half membrane-spanning helices around the channel. This structure elucidates the mechanism of selective permeability for linear carbohydrates and suggests how ions and water are excluded.
- Department of Biochemistry and Biophysics, School of Medicine, University of California, San Francisco, CA 94143-0448, USA.
Organizational Affiliation: