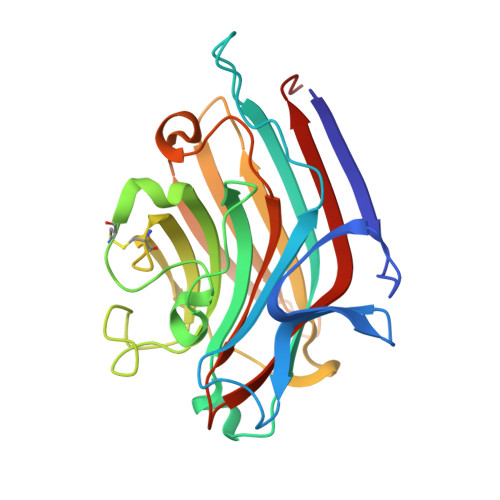
The 2.2 A resolution structure of the O(H) blood-group-specific lectin I from Ulex europaeus.
Audette, G.F., Vandonselaar, M., Delbaere, L.T.(2000) J Mol Biology 304: 423-433
- PubMed: 11090284 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4214
- Primary Citation Related Structures:
1FX5 - PubMed Abstract:
The tertiary and quaternary structure of the lectin I from Ulex europaeus (UE-I) has been determined to 2.2 A resolution. UE-I is a dimeric metalloglycoprotein that binds the H-type 2 human blood group determinant [alpha-L-Fucalpha(1-->2)-beta-D-Galbeta(1-->4)-beta-D-Glc NAcalpha-]. Nine changes from the published amino acid sequence were necessary to account for the electron density. The quaternary structural organization of UE-I is that of the most commonly occurring legume lectin dimer. The tertiary structure of the monomeric subunits is similar to that in the conventional lectin subunit; however, some structural differences are noted. These differences include a four-stranded anti-parallel "S" sheet in UE-I versus the five-stranded S sheet in other lectin monomers. The Ala residue of the Ala-Asp cis-peptide bond present in the carbohydrate-binding site of the conventional lectin monomer is replaced with a Thr in the UE-I structure. Also, a novel disulfide bridge linking Cys115 and Cys150 is present. There are two metallic ions, one calcium and the other manganese, per subunit. N-linked oligosaccharides are at residues 23 and 111 of each subunit. One molecule of R-2-methyl-2, 4-pentanediol (R-MPD) is present in a shallow depression on the surface of each subunit. In order to examine the binding of the H-type 2 blood group determinant by UE-I, its beta-methyl glycoside (H-type 2-OMe) was docked into the binding site of R-MPD. The epitope previously identified for H-type 2-OMe by chemical mapping proved, with only minor adjustment of amino acid residues, to be complementary to the shallow cavity occupied by R-MPD in the structure. Several key interactions have been proposed between the H-type 2-OMe and UE-I.
- Department of Biochemistry, University of Saskatchewan, 107 Wiggins Road, Saskatoon, Saskatchewan, S7N 5E5, Canada.
Organizational Affiliation: