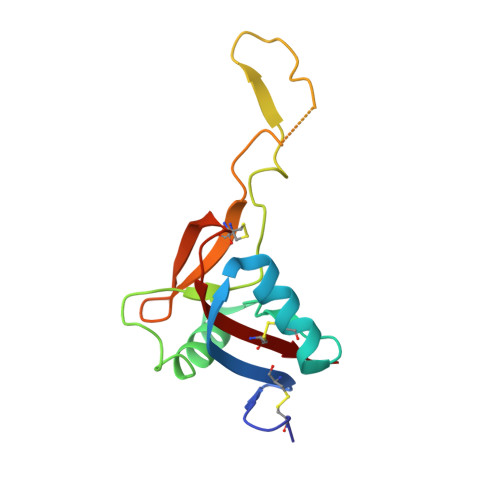
Crystal structure of the von Willebrand factor modulator botrocetin.
Sen, U., Vasudevan, S., Subbarao, G., McClintock, R.A., Celikel, R., Ruggeri, Z.M., Varughese, K.I.(2001) Biochemistry 40: 345-352
- PubMed: 11148028 Search on PubMed
- DOI: https://doi.org/10.1021/bi0021737
- Primary Citation Related Structures:
1FVU - PubMed Abstract:
The binding of von Willebrand factor (vWF) to the platelet receptor, glycoprotein (GP) Ib-IX-V complex, has a key role in the initiation of thrombus formation and is regulated by interactions with extracellular matrix components under the influence of hemodynamic forces. To a certain extent, these effects can be mimicked in vitro by two nonphysiologic modulators, ristocetin and botrocetin. The latter, isolated from the venom of the snake Bothrops jararaca, is a 31-kDa heterodimeric protein that forms a soluble complex with vWF. As an initial step toward understanding the mechanisms that regulate vWF function, we have solved the crystal structure of botrocetin at 1.8 A resolution. Botrocetin exhibits homology with other snake proteins, but contains only one metal binding site as compared to two in Factor IX binding protein and Factor IX/X binding protein and none in flavocetin. A distinctive feature of botrocetin is the presence of a negatively charged surface that may play a role in the association with the vWF A1 domain.
- Roon Research Center for Arteriosclerosis and Thrombosis, Division of Experimental Hemostasis and Thrombosis, Department of Molecular and Experimental Medicine, The Scripps Research Institute, La Jolla, California 92037, USA.
Organizational Affiliation: