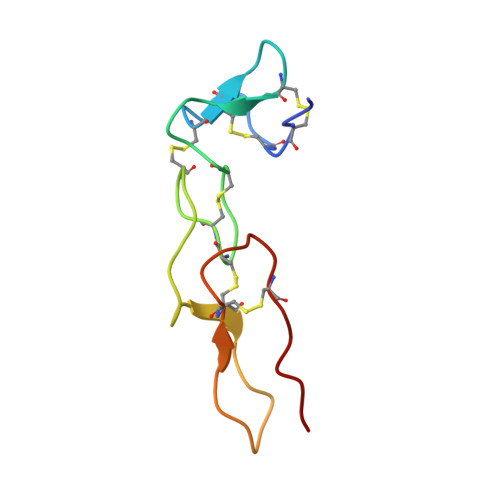
The nuclear magnetic resonance solution structure of flavoridin, an antagonist of the platelet GP IIb-IIIa receptor.
Senn, H., Klaus, W.(1993) J Mol Biology 232: 907-925
- PubMed: 8355277
- DOI: https://doi.org/10.1006/jmbi.1993.1439
- Primary Citation of Related Structures:
1FVL - PubMed Abstract:
The snake venom protein flavoridin, a polypeptide of 70 amino acid residues, is a potent inhibitor of blood platelet aggregation. It binds to cell-surface integrin receptors such as the fibrinogen receptor glycoprotein IIb/IIIa. The inhibitory properties of flavoridin have been attributed to the tripeptide segment Arg-Gly-Asp (residues 49 to 51). This paper describes the determination of the three-dimensional structure of flavoridin in aqueous solution based on two-dimensional nuclear magnetic resonance spectroscopy. A family of 18 conformers was selected to characterize the solution structure. The molecule comprises two structural domains, an N-terminal unit extending from residues 1 to 25, and a C-terminal unit from residues 26 to 70. Whereas the mutual spatial orientation of these regions is not well defined, each one is well organized within itself. The segment 26 to 70, which is homologous to the sequence of the snake toxins echistatin and eristostatin, shows an average value of 1.0 A for the root-mean-square deviations of the backbone atoms among the 18 conformers. The structure of flavoridin consists essentially of non-repetitive elements such as tight turns and loops, whose location and conformation are characterized in this paper. With the exception of two short regions of antiparallel beta-sheet, no classic element of protein secondary structure is present. The six disulphide bridges, which have been mapped by applying a novel computational strategy (see accompanying paper), are the dominant organizational feature of the polypeptide fold of flavoridin. Two of the bridges are located in the N-terminal domain, three in the C-terminal domain and one connects the two structural units. The mobile RGD recognition sequence for integrins is located peripheral to the core region of the C-terminal domain at the most exposed end of a nine residue loop structure, which is attached to a short beta-sheet. The C terminus is close to this loop structure.
- Department of Pharmaceutical Research-New Technologies F. Hoffmann-LaRoche Ltd, Basel, Switzerland.
Organizational Affiliation: