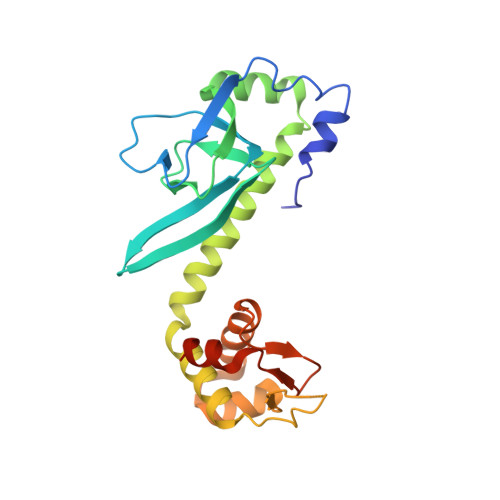
Structure of the CO sensing transcription activator CooA.
Lanzilotta, W.N., Schuller, D.J., Thorsteinsson, M.V., Kerby, R.L., Roberts, G.P., Poulos, T.L.(2000) Nat Struct Biol 7: 876-880
- PubMed: 11017196 Search on PubMed
- DOI: https://doi.org/10.1038/82820
- Primary Citation Related Structures:
1FT9 - PubMed Abstract:
CooA is a homodimeric transcription factor that belongs to the catabolite activator protein (CAP) family. Binding of CO to the heme groups of CooA leads to the transcription of genes involved in CO oxidation in Rhodospirillum rubrum. The 2.6 A structure of reduced (Fe2+) CooA reveals that His 77 in both subunits provides one heme ligand while the N-terminal nitrogen of Pro 2 from the opposite subunit provides the other ligand. A structural comparison of CooA in the absence of effector and DNA (off state) with that of CAP in the effector and DNA bound state (on state) leads to a plausible model for the mechanism of allosteric control in this class of proteins as well as the CO dependent activation of CooA.
- Departments of Molecular Biology and Biochemistry and Physiology and Biophysics and the Program in Macromolecular Structure, University of California, Irvine, California 92697-3900, USA.
Organizational Affiliation: