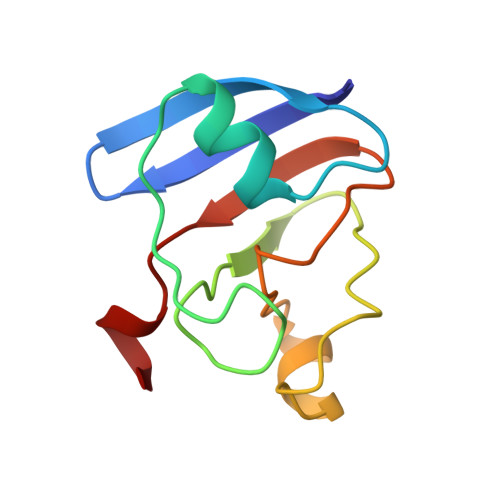
Structure of [2Fe-2S] ferredoxin I from Equisetum arvense at 1.8 A resolution.
Ikemizu, S., Bando, M., Sato, T., Morimoto, Y., Tsukihara, T., Fukuyama, K.(1994) Acta Crystallogr D Biol Crystallogr 50: 167-174
- PubMed: 15299454 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444993009588
- Primary Citation Related Structures:
1FRR - PubMed Abstract:
Ferredoxin I (Fd I) from Equisetum arvense is an iron-sulfur protein composed of 95 amino-acid residues and one [2Fe-2S] cluster. It crystallized in the space group P2(1), a = 30.4, b = 57.4, c = 47.5 A and beta = 78.7 degrees with two molecules per asymmetric unit. X-ray diffraction data up to 1.8 A resolution were collected by using a Rigaku four-circle diffractometer. The initial model of Fd I, which was derived by the molecular replacement method using a structure of the Fd I from the blue-green alga Aphanothece sacrum, was refined by molecular dynamics simulation and a least-squares minimization with stereochemical restraints. Positional parameters and isotropic temperature factors for 1420 non-H protein atoms and 183 water molecules were refined on 13 838 observed structure factors (F(o) > sigma(Fo)) between 10.0 and 1.8 A resolution. The final Rfactor was 17.0%, and the standard deviation of atomic position estimated by Luzzati plot [Luzzati (1952). Acta Cryst. 5, 802-810] was 0.2 A. The electron-density map was well defined for the two independent molecules except for the N-terminal residue and the three C-terminal residues. Equivalent Calpha atoms of two independent molecules in the asymmetric unit were superposed by the least-squares method with root-mean-square deviations of 0.26 A. Reasonable structural differences were observed at a polypeptide segment having few intramolecular interactions. Highly flexible regions of the molecule were assigned from the structural differences between the two independent molecules in the crystal and the distribution of temperature factors along the polypeptide chain.
- The Graduate University for Advanced Studies, Oho, Tsukuba, Ibaraki, Japan.
Organizational Affiliation: