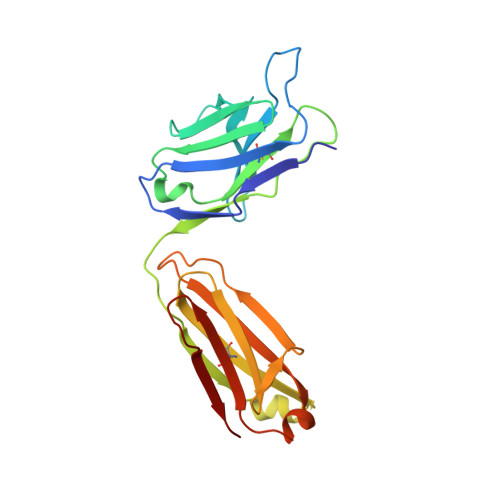
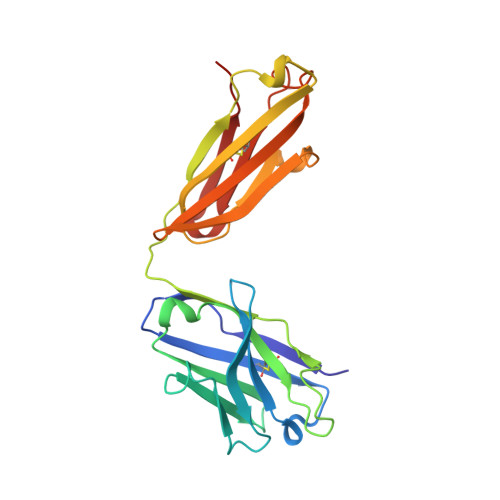
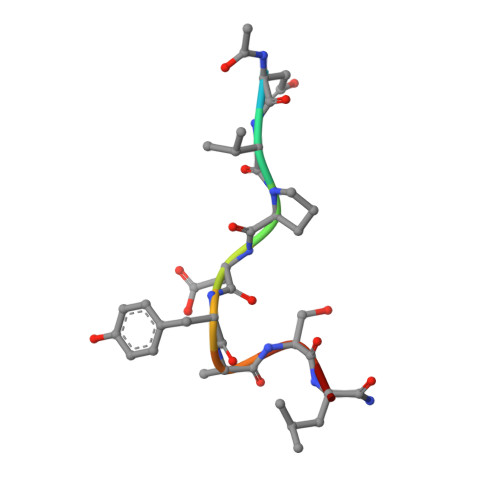
Crystal structure of a peptide complex of anti-influenza peptide antibody Fab 26/9. Comparison of two different antibodies bound to the same peptide antigen.
Churchill, M.E., Stura, E.A., Pinilla, C., Appel, J.R., Houghten, R.A., Kono, D.H., Balderas, R.S., Fieser, G.G., Schulze-Gahmen, U., Wilson, I.A.(1994) J Mol Biology 241: 534-556
- PubMed: 7520084 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1530
- Primary Citation Related Structures:
1FRG - PubMed Abstract:
The three-dimensional structure of the complex of a second anti-peptide antibody (Fab 26/9) that recognizes the same six-residue epitope of an immunogenic peptide from influenza virus hemagglutinin (HA1; 75-110) as Fab 17/9 with the peptide has been determined at 2.8 A resolution. The amino acid sequence of the variable region of the 26/9 antibody differs in 24 positions from that of 17/9, the first antibody in this series for which several ligand-bound and free structures have been determined and refined. Comparison of the 26/9-peptide with the 17/9-peptide complex structures shows that the two Fabs are very similar (r.m.s.d. 0.5 to 0.8 A) and that the peptide antigen (101-107) has virtually the same conformation (r.m.s.d. 0.3 to 0.8 A) when bound to both antibodies. A sequence difference in the 26/9 binding pocket (L94; His in 26/9, Asn in 17/9) results in an interaction with a bound water molecule that is not seen in the 17/9 structures. Epitope mapping shows that the relative specificity of 26/9 and 17/9 antibodies for individual positions of the peptide antigen are slightly different. Amino acid substitutions in the peptide, particularly at position SerP107, are tolerated to different extents by 17/9 and 26/9. Structural and sequence analysis suggests that amino acid differences near the peptide-binding site are responsible for altering slightly the specificity of 26/9 for three peptide residues and illustrates how amino acid substitutions can modify antibody-antigen interactions and thereby modulate antibody specificity.
- Department of Molecular Biology, Scripps Research Institute, La Jolla, CA 92037.
Organizational Affiliation: