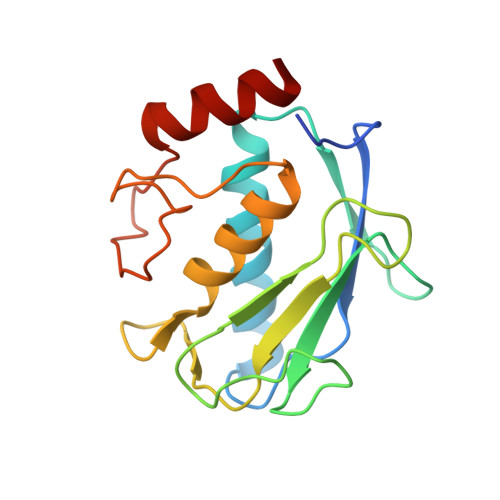
High-resolution solution structure of the catalytic fragment of human collagenase-3 (MMP-13) complexed with a hydroxamic acid inhibitor.
Moy, F.J., Chanda, P.K., Chen, J.M., Cosmi, S., Edris, W., Levin, J.I., Powers, R.(2000) J Mol Biology 302: 671-689
- PubMed: 10986126 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.4082
- Primary Citation Related Structures:
1FLS, 1FM1 - PubMed Abstract:
The high-resolution solution structure of the catalytic fragment of human collagenase-3 (MMP-13) complexed with a sulfonamide derivative of a hydroxamic acid compound (WAY-151693) has been determined by multidimensional heteronuclear NMR. A total of 30 structures were calculated for residues 7-164 by means of hybrid distance geometry-simulated annealing using a total of 3280 experimental NMR restraints. The atomic rms distribution about the mean coordinate positions for the 30 structures is 0.43(+/-0.05) A for the backbone atoms, 0.80(+/-0.09) A for all atoms, and 0.47(+/-0.04) A for all atoms excluding disordered side-chains. The overall structure of MMP-13 is composed of a beta-sheet consisting of five beta-strands in a mixed parallel and anti-parallel arrangement and three alpha-helices where its overall fold is consistent with previously solved MMP structures. A comparison of the NMR structure of MMP-13 with the published 1.6 A resolution X-ray structure indicates that the major differences between the structures is associated with loop dynamics and crystal-packing interactions. The side-chains of some active-site residues for the NMR and X-ray structures of MMP-13 adopt distinct conformations. This is attributed to the presence of unique inhibitors in the two structures that encounter distinct interactions with MMP-13. The major structural difference observed between the MMP-13 and MMP-1 NMR structures is the relative size and shape of the S1' pocket where this pocket is significantly longer for MMP-13, nearly reaching the surface of the protein. Additionally, MMP-1 and MMP-13 exhibit different dynamic properties for the active-site loop and the structural Zn-binding region. The inhibitor WAY-151693 is well defined in the MMP-13 active-site based on a total of 52 distance restraints. The binding motif of WAY-151693 in the MMP-13 complex is consistent with our previously reported MMP-1:CGS-27023A NMR structure and is similar to the MMP-13: RS-130830 X-ray structure.
- Department of Biological Chemistry, Wyeth Research, 85 Bolton St., Cambridge, MA 02140, USA.
Organizational Affiliation: