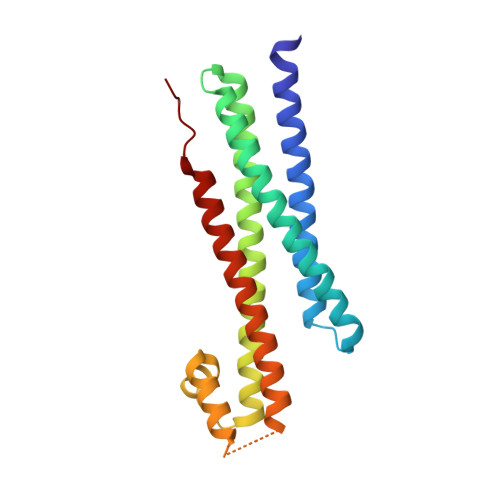
Interactions within the yeast t-SNARE Sso1p that control SNARE complex assembly.
Munson, M., Chen, X., Cocina, A.E., Schultz, S.M., Hughson, F.M.(2000) Nat Struct Biol 7: 894-902
- PubMed: 11017200 Search on PubMed
- DOI: https://doi.org/10.1038/79659
- Primary Citation Related Structures:
1FIO - PubMed Abstract:
In the eukaryotic secretory and endocytic pathways, transport vesicles shuttle cargo among intracellular organelles and to and from the plasma membrane. Cargo delivery entails fusion of the transport vesicle with its target, a process thought to be mediated by membrane bridging SNARE protein complexes. Temporal and spatial control of intracellular trafficking depends in part on regulating the assembly of these complexes. In vitro, SNARE assembly is inhibited by the closed conformation adopted by the syntaxin family of SNAREs. To visualize this closed conformation directly, the X-ray crystal structure of a yeast syntaxin, Sso1p, has been determined and refined to 2.1 A resolution. Mutants designed to destabilize the closed conformation exhibit accelerated rates of SNARE assembly. Our results provide insight into the mechanism of SNARE assembly and its intramolecular and intermolecular regulation.
- Department of Molecular Biology, Princeton University, Princeton, New Jersey 08544, USA.
Organizational Affiliation: