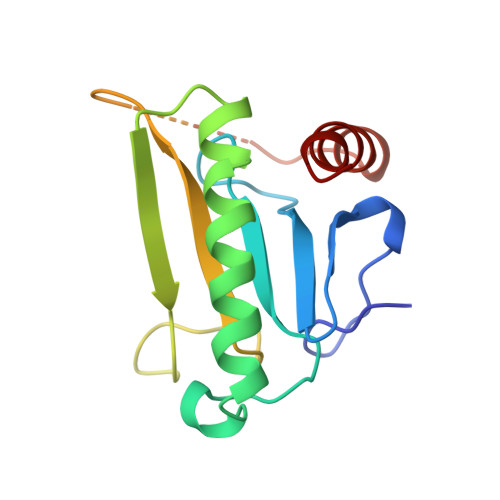
Genetic, biochemical, and crystallographic characterization of Fhit-substrate complexes as the active signaling form of Fhit.
Pace, H.C., Garrison, P.N., Robinson, A.K., Barnes, L.D., Draganescu, A., Rosler, A., Blackburn, G.M., Siprashvili, Z., Croce, C.M., Huebner, K., Brenner, C.(1998) Proc Natl Acad Sci U S A 95: 5484-5489
- PubMed: 9576908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.95.10.5484
- Primary Citation Related Structures:
1FHI, 2FHI - PubMed Abstract:
Alterations in the FHIT gene at 3p14.2 occur as early and frequent events in the development of several common human cancers. The ability of human Fhit-negative cells to form tumors in nude mice is suppressed by stable reexpression of Fhit protein. Fhit protein is a diadenosine P1,P3-triphosphate (ApppA) hydrolase whose fungal and animal homologs form a branch of the histidine triad (HIT) superfamily of nucleotide-binding proteins. Because the His-96 --> Asn substitution of Fhit, which retards ApppA hydrolase activity by seven orders of magnitude, did not block tumor-suppressor activity in vivo, we determined whether this mutation affected ApppA binding or particular steps in the ApppA catalytic cycle. Evidence is presented that His-96 --> Asn protein binds ApppA well and forms an enzyme-AMP intermediate extremely poorly, suggesting that Fhit-substrate complexes are the likely signaling form of the enzyme. The cocrystal structure of Fhit bound to Ado-p-CH2-p-ps-Ado (IB2), a nonhydrolyzable ApppA analog, was refined to 3.1 A, and the structure of His-96 --> Asn Fhit with IB2 was refined to 2.6 A, revealing that two ApppA molecules bind per Fhit dimer; identifying two additional adenosine-binding sites on the dimer surface; and illustrating that His-98 is positioned to donate a hydrogen bond to the scissile bridging oxygen of ApppA substrates. The form of Fhit bound to two ApppA substrates would present to the cell a dramatically phosphorylated surface, prominently displaying six phosphate groups and two adenosine moieties in place of a deep cavity lined with histidines, arginines, and glutamines.
- Kimmel Cancer Institute, Thomas Jefferson University, Philadelphia, PA 19107, USA.
Organizational Affiliation: