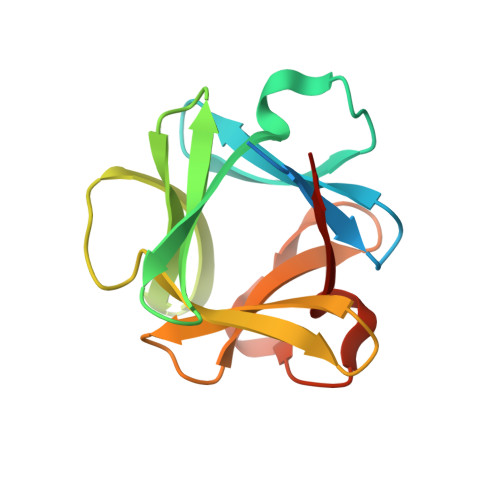
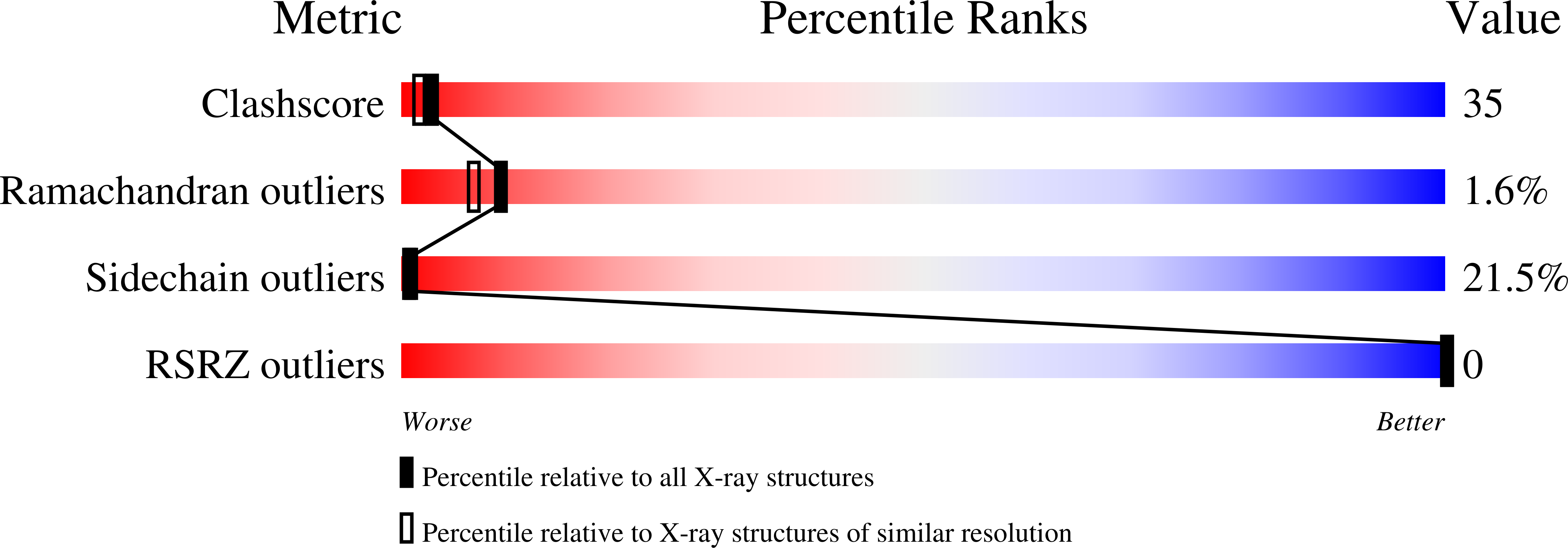Refinement of the structure of human basic fibroblast growth factor at 1.6 A resolution and analysis of presumed heparin binding sites by selenate substitution.
Eriksson, A.E., Cousens, L.S., Matthews, B.W.(1993) Protein Sci 2: 1274-1284
- PubMed: 7691311 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560020810
- Primary Citation Related Structures:
1FGA, 4FGF - PubMed Abstract:
The three-dimensional structure of human basic fibroblast growth factor has been refined to a crystallographic residual of 16.1% at 1.6 A resolution. The structure has a Kunitz-type fold and is composed of 12 antiparallel beta-strands, 6 of which form a beta-barrel. One bound sulfate ion has been identified in the model, hydrogen bonded to the side chains of Asn 27, Arg 120, and Lys 125. The side chain of Arg 120 has two conformations, both of which permit hydrogen bonds to the sulfate. This sulfate binding site has been suggested as the binding site for heparin (Eriksson, A.E., Cousens, L.S., Weaver, L.H., & Matthews, B.W., 1991, Proc. Natl. Acad. Sci. USA 88, 3441-3445). Two beta-mercaptoethanol (BME) molecules are also included in the model, each forming a disulfide bond to the S gamma atoms of Cys 69 and Cys 92, respectively. The side chain of Cys 92 has two conformations of which only one can bind BME. Therefore the BME molecule is half occupied at this site. The locations of possible sulfate binding sites on the protein were examined by replacing the ammonium sulfate in the crystallization medium with ammonium selenate. Diffraction data were measured to 2.2 A resolution and the structure refined to an R-factor of 13.8%. The binding of the more electron-dense selenate ion was identified at two positions. One position was identical to the sulfate binding site identified previously. The second selenate binding site, which is of lower occupancy, is situated 5.6 A from the first. This ion is hydrogen bonded by the side chain of Lys 135 and Arg 120. Thus the side chain of Arg 120 binds two selenate ions simultaneously. It is suggested that the observed second selenate binding site should also be considered as a possible binding site for heparin, or that both selenate binding sites might simultaneously contribute to the binding of heparin.
- Institute of Molecular Biology, Howard Hughes Medical Institute, University of Oregon, Eugene 97403.
Organizational Affiliation: