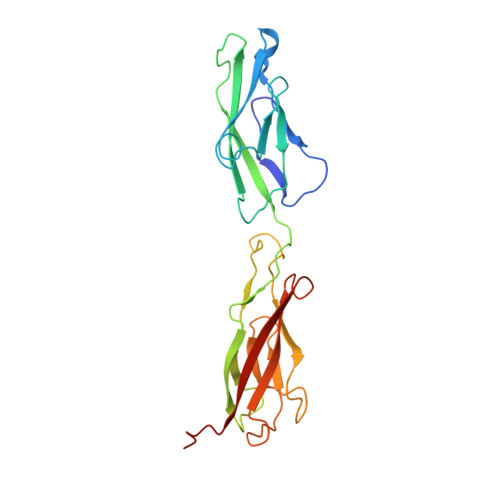
A new crystal structure, Ca2+ dependence and mutational analysis reveal molecular details of E-cadherin homoassociation.
Pertz, O., Bozic, D., Koch, A.W., Fauser, C., Brancaccio, A., Engel, J.(1999) EMBO J 18: 1738-1747
- PubMed: 10202138 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.7.1738
- Primary Citation Related Structures:
1FF5 - PubMed Abstract:
Electron microscopy of ECADCOMP, a recombinant E-cadherin ectodomain pentamerized by the assembly domain of cartilage oligomeric matrix protein, has been used to analyze the role of cis-dimerization and trans-interaction in the homophilic association of this cell adhesion molecule. The Ca2+ dependency of both interactions was investigated. Low Ca2+ concentrations (50 microM) stabilized the rod-like structure of E-cadherin. At medium Ca2+ concentration (500 microM), two adjacent ectodomains in a pentamer formed cis-dimers. At high Ca2+ concentration (>1 mM), two cis-dimers from different pentamers formed a trans-interaction. The X-ray structure of an N-terminal domain pair of E-cadherin revealed two molecules per asymmetric unit in an intertwisted X-shaped arrangement with closest contacts in the Ca2+-binding region between domains 1 and 2. Contrary to previous data, Trp2 was docked in the hydrophobic cavity of its own molecule, and was therefore not involved in cis-dimerization of two molecules. This was supported further by W2A and A80I (a residue involved in the hydrophobic cavity surrounding Trp2) mutations in ECADCOMP which both led to abrogation of the trans- but not the cis-interaction. Structural and biochemical data suggest a link between Ca2+ binding in the millimolar range and Trp2 docking, both events being essential for the trans-association.
- Department of Biophysical Chemistry, Biozentrum, University of Basel, Klingelbergstrasse 70, 4056 Basel, Switzerland.
Organizational Affiliation: