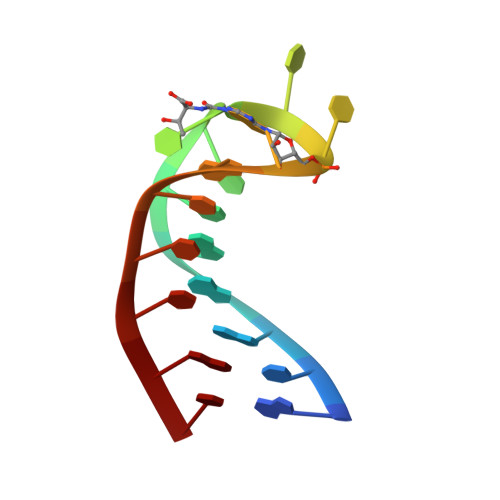
Functional anticodon architecture of human tRNALys3 includes disruption of intraloop hydrogen bonding by the naturally occurring amino acid modification, t6A.
Stuart, J.W., Gdaniec, Z., Guenther, R., Marszalek, M., Sochacka, E., Malkiewicz, A., Agris, P.F.(2000) Biochemistry 39: 13396-13404
- PubMed: 11063577 Search on PubMed
- DOI: https://doi.org/10.1021/bi0013039
- Primary Citation Related Structures:
1FEQ - PubMed Abstract:
The structure of the human tRNA(Lys3) anticodon stem and loop domain (ASL(Lys3)) provides evidence of the physicochemical contributions of N6-threonylcarbamoyladenosine (t(6)A(37)) to tRNA(Lys3) functions. The t(6)A(37)-modified anticodon stem and loop domain of tRNA(Lys3)(UUU) (ASL(Lys3)(UUU)- t(6)A(37)) with a UUU anticodon is bound by the appropriately programmed ribosomes, but the unmodified ASL(Lys3)(UUU) is not [Yarian, C., Marszalek, M., Sochacka, E., Malkiewicz, A., Guenther, R., Miskiewicz, A., and Agris, P. F., Biochemistry 39, 13390-13395]. The structure, determined to an average rmsd of 1.57 +/- 0.33 A (relative to the mean structure) by NMR spectroscopy and restrained molecular dynamics, is the first reported of an RNA in which a naturally occurring hypermodified nucleoside was introduced by automated chemical synthesis. The ASL(Lys3)(UUU)-t(6)A(37) loop is significantly different than that of the unmodified ASL(Lys3)(UUU), although the five canonical base pairs of both ASL(Lys3)(UUU) stems are in the standard A-form of helical RNA. t(6)A(37), 3'-adjacent to the anticodon, adopts the form of a tricyclic nucleoside with an intraresidue H-bond and enhances base stacking on the 3'-side of the anticodon loop. Critically important to ribosome binding, incorporation of the modification negates formation of an intraloop U(33).A(37) base pair that is observed in the unmodified ASL(Lys3)(UUU). The anticodon wobble position U(34) nucleobase in ASL(Lys3)(UUU)-t(6)A(37) is significantly displaced from its position in the unmodified ASL and directed away from the codon-binding face of the loop resulting in only two anticodon bases for codon binding. This conformation is one explanation for ASL(Lys3)(UUU) tendency to prematurely terminate translation and -1 frame shift. At the pH 5.6 conditions of our structure determination, A(38) is protonated and positively charged in ASL(Lys3)(UUU)-t(6)A(37) and the unmodified ASL(Lys3)(UUU). The ionized carboxylic acid moiety of t(6)A(37) possibly neutralizes the positive charge of A(+)(38). The protonated A(+)(38) can base pair with C(32), but t(6)A(37) may weaken the interaction through steric interference. From these results, we conclude that ribosome binding cannot simply be an induced fit of the anticodon stem and loop, otherwise the unmodified ASL(Lys3)(UUU) would bind as well as ASL(Lys3)(UUU)-t(6)A(37). t(6)A(37) and other position 37 modifications produce the open, structured loop required for ribosomal binding.
- Department of Biochemistry, North Carolina State University, Raleigh, North Carolina 27695-7622, USA.
Organizational Affiliation: