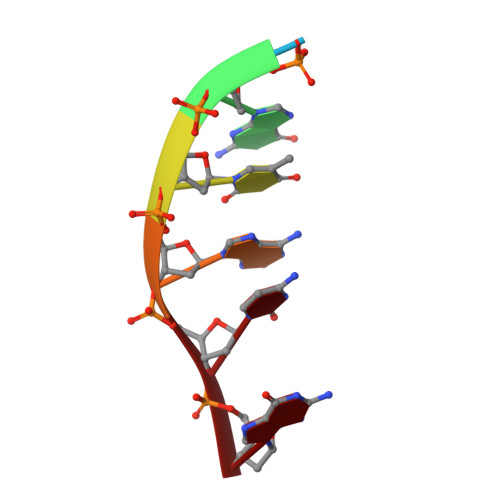
Binding of a macrocyclic bisacridine and ametantrone to CGTACG involves similar unusual intercalation platforms.
Yang, X.L., Robinson, H., Gao, Y.G., Wang, A.H.(2000) Biochemistry 39: 10950-10957
- PubMed: 10998231 Search on PubMed
- Primary Citation Related Structures:
1FD5, 1FDG - PubMed Abstract:
The binding of a macrocyclic bisacridine and an antitumor intercalator ametantrone to DNA has been studied. We carried out X-ray diffraction analyses of the complexes between both intercalators and CGTACG. We have determined the crystal structure, by the multiple-wavelength anomalous diffraction (MAD) method, of bisacridine complexed with CGTA[br(5)C]G at 1.8 A resolution. The refined native crystal structure at 1.1 A resolution (space group C222, a = 29.58 A, b = 54.04 A, c = 40.22 A, and R-factor = 0.163) revealed that only one acridine of the bisacridine drug binds at the C5pG6 step of the DNA, with the other acridine plus both linkers completely disordered. Surprisingly, both terminal G.C base pairs are unraveled. The C1 nucleotide is disordered, and the G2 base is bridged to its own phosphate P2 through a hydrated Co(2+) ion. G12 is swung toward the minor groove with its base stacked over the backbone. The C7 nucleotide is flipped away from the duplex part and base paired to a 2-fold symmetry-related G6. The central four base pairs adopt the B-DNA conformation. An unusual intercalator platform is formed by bringing four complexes together (involving the 222 symmetry) such that the intercalator cavity is flanked by two sets of G x C base pairs (i.e., C5 x G8 and G6 x C7) on each side, joined together by G6 x G8 tertiary base pairing interactions. In the bisacridine-CGTACG complex, the intercalation platform is intercalated with two acridines, whereas in the ametantrone-CGTACG complex, only one ametantrone is bound. NMR titration of the bisacridine to AACGATCGTT suggests that the bisacridine prefers to bridge more than one DNA duplex by intercalating each acridine to different duplexes. The results may be relevant in understanding binding of certain intercalators to DNA structure associated with the quadruplet helix and Holliday junction.
- Department of Biochemistry, School of Molecular & Cellular Biology, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801, USA.
Organizational Affiliation: