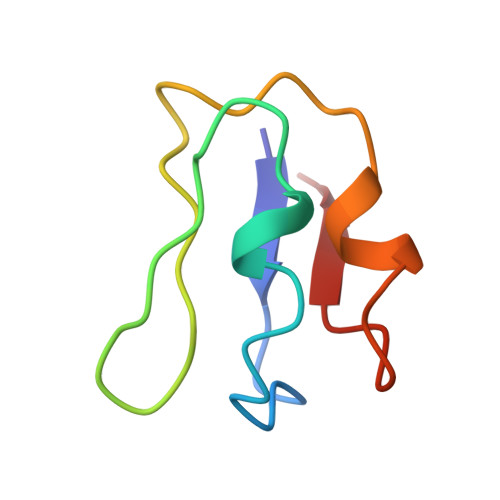
Structure of the ferredoxin from Clostridium acidurici: model at 1.8 A resolution.
TranQui, D., Jesior, J.C.(1995) Acta Crystallogr D Biol Crystallogr 51: 155-159
- PubMed: 15299316 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444994010735
- Primary Citation Related Structures:
1FCA - PubMed Abstract:
Ferredoxins (Fd) are electron-carrier proteins, the active sites of which are organized around clusters made of iron and inorganic sulfur. The Fd from Clostridium acidurici is 55 amino acids long and contains two [4Fe-4S] clusters. Crystals have been obtained in the space group P4(3)2(1)2, a = b = 34.441 (5), c = 74.778 (9) A. The structure was solved by molecular replacement using the Fd from Peptostreptcoccus asaccharolyticus as a search model, these two ferredoxins having 37 residues in common. Refinement using molecular-dynamics techniques was then initiated. Successive rounds of model building and refinement gave a structure that includes 45 water molecules with R = 15%. At this stage, the electron-density map clearly revealed discrepancies in the position of two amino acids in the published primary sequence. Refinement based on these modifications led to R = 14.3% for 3921 reflections up to 1.8 A, resolution. The geometry of the two clusters has been found to be in good agreement with that previously obtained at a lower resolution. Interactions of polypeptide chain with the [4Fe-4S] clusters, the cluster geometry as well as the hydrogen bonds involving S, Sgamma, N and water molecules are reported.
- Laboratoire de Cristallographie, CNRS, Grenoble, France.
Organizational Affiliation: