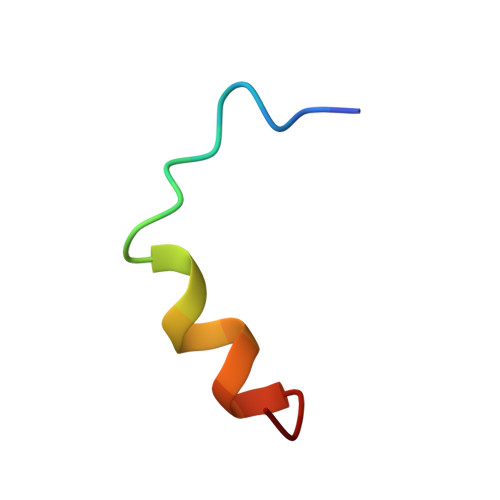
Structure and topography of the membrane-binding C2 domain of factor VIII in the presence of dodecylphosphocholine micelles.
Veeraraghavan, S., Baleja, J.D., Gilbert, G.E.(1998) Biochem J 332: 549-555
- PubMed: 9601086
- DOI: https://doi.org/10.1042/bj3320549
- Primary Citation of Related Structures:
1FAC - PubMed Abstract:
A 21 residue peptide from the C2 domain of the antihaemophilic factor VIII competes with factor VIII for membrane-binding sites in vitro. Here, we provide the structure and topography of the peptide in solution, on dodecylphosphocholine (DPC) micelles, determined using 1H-NMR spectroscopy. The peptide assumes an amphipathic structure comprising an extended N-terminal region and a C-terminal helix. The average root-mean-square deviation is 0.7+/-0.2 A for the superimposition of the backbone atoms of Ile6 to Arg18 on the lowest energy structure. Whereas the backbone conformation is similar to that in SDS micelles, the Trp11 side-chain orientation is dramatically changed. The indole ring is nearly parallel to the peptide backbone in SDS micelles but perpendicular in DPC micelles. Further, pKa values of the two histidines change by more than 1 pH unit in SDS relative to DPC, which localizes the imidazole rings to the interfacial region. Line-broadening induced by spin-labelled phosphatidylcholine shows that most of the amino acid side-chains that penetrate the DPC micelle are hydrophobic. Thus, the long axis of the peptide lies parallel to the micelle surface and the hydrophobic face of the alpha-helix provides hydrophobic membrane interaction. The large chemical shift changes shown by Trp11 and N-terminal amino acid residues in SDS relative to DPC indicate that this region may be involved in membrane phospholipid recognition. 1H-NMR assignments, CD spectra, one-dimensional 1H-NMR spectra, chemical-shift analysis and nuclear Overhauser effect information are reported in Supplementary Publication SUP 50184 (11 pages), which has been deposited at the British Library Document Supply Centre, Boston Spa, Wetherby, West Yorkshire LS23 7BQ, U.K, from whom copies can be obtained according to the terms indicated in Biochem. J. (1997) 321, 8.
- Department of Biochemistry, Tufts University School of Medicine, 136 Harrison Ave., Boston, MA 02111, USA.
Organizational Affiliation: