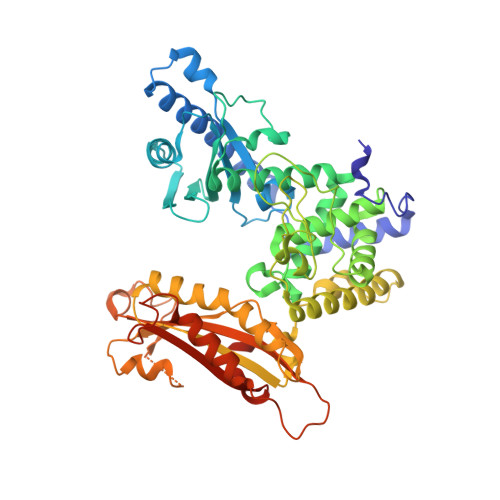
Structure of yeast poly(A) polymerase alone and in complex with 3'-dATP.
Bard, J., Zhelkovsky, A.M., Helmling, S., Earnest, T.N., Moore, C.L., Bohm, A.(2000) Science 289: 1346-1349
- PubMed: 10958780 Search on PubMed
- DOI: https://doi.org/10.1126/science.289.5483.1346
- Primary Citation Related Structures:
1FA0 - PubMed Abstract:
Polyadenylate [poly(A)] polymerase (PAP) catalyzes the addition of a polyadenosine tail to almost all eukaryotic messenger RNAs (mRNAs). The crystal structure of the PAP from Saccharomyces cerevisiae (Pap1) has been solved to 2.6 angstroms, both alone and in complex with 3'-deoxyadenosine triphosphate (3'-dATP). Like other nucleic acid polymerases, Pap1 is composed of three domains that encircle the active site. The arrangement of these domains, however, is quite different from that seen in polymerases that use a template to select and position their incoming nucleotides. The first two domains are functionally analogous to polymerase palm and fingers domains. The third domain is attached to the fingers domain and is known to interact with the single-stranded RNA primer. In the nucleotide complex, two molecules of 3'-dATP are bound to Pap1. One occupies the position of the incoming base, prior to its addition to the mRNA chain. The other is believed to occupy the position of the 3' end of the mRNA primer.
- Boston Biomedical Research Institute, 64 Grove Street, Watertown, MA 02472, USA.
Organizational Affiliation: