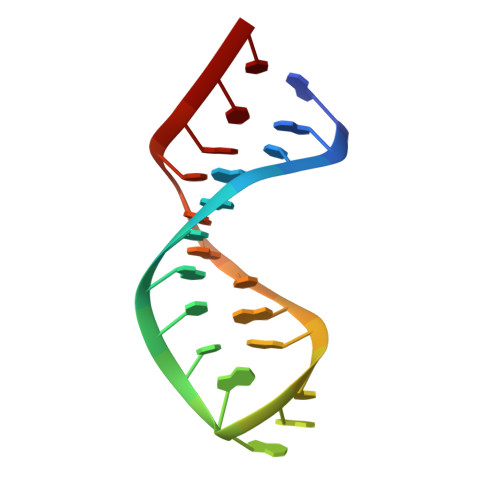
Solution structure of Cobalt(III)hexammine complexed to the GAAA tetraloop, and metal-ion binding to G.A mismatches.
Rudisser, S., Tinoco Jr., I.(2000) J Mol Biology 295: 1211-1232
- PubMed: 10653698 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.3421
- Primary Citation Related Structures:
1F9L - PubMed Abstract:
The solution structure of a 22 nt RNA hairpin and its complex with Co(NH(3))(6)(3+) bound to the GAAA tetraloop has been determined by NMR spectroscopy. Co(NH(3))(6)(3+) has a similar geometry to Mg(H(2)O)(6)(2+) and can be used as a probe for binding sites of completely solvated magnesium ions. The hairpin contains tandem G.A mismatches, similar to the P5abc region of a group I intron, and is closed by a GAAA tetraloop. The tandem G.A mismatches are imino hydrogen bonded in contrast with the sheared G.A mismatches found in a different context in the crystal structure of the P4-P6 domain. Chemical shift changes of the imino protons upon titration of the RNA hairpin with Mg(2+) and with Co(NH(3))(6)(3+) were used to identify ion-binding sites. Paramagnetic resonance broadening upon titration with Mn(2+) was also used. The titration curves gave dissociation binding constants for the magnesium ions in the millimolar range, similar to the binding in the major groove of RNA at tandem G.U base-pairs. Although the largest chemical shift change occurred at an imino proton of one of the G.A base-pairs, no nuclear Overhauser enhancement cross-peaks between the cobalt ligand and neighboring RNA protons were seen, presumably due to the high mobility of the Co(NH(3))(6)(3+) at this site. Nuclear Overhauser enhancement cross-peaks between Co(NH(3))(6)(3+) and the GAAA tetraloop were observed, which allowed the determination of the structure of the tetraloop binding site. The Co(NH(3))(6)(3+) is bound in the major groove of the GAAA tetraloop with hydrogen bonds to guanine base N7 and to phosphate oxygen atoms of the tetraloop.
- Department of Chemistry, University of California and Physical Biosciences Division Lawrence Berkeley National Laboratory, Berkeley, CA, 94720-1460, USA.
Organizational Affiliation: