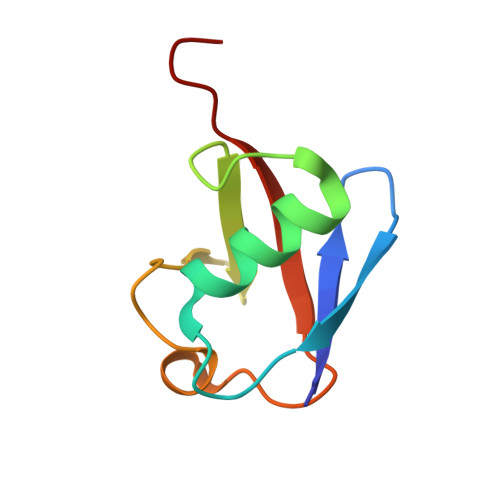
Structure of a new crystal form of tetraubiquitin.
Phillips, C.L., Thrower, J., Pickart, C.M., Hill, C.P.(2001) Acta Crystallogr D Biol Crystallogr 57: 341-344
- PubMed: 11173499 Search on PubMed
- DOI: https://doi.org/10.1107/s090744490001800x
- Primary Citation Related Structures:
1F9J - PubMed Abstract:
Polyubiquitin chains, in which the C-terminus and a lysine side chain of successive ubiquitin molecules are linked by an isopeptide bond, function to target substrate proteins for degradation by the 26S proteasome. Chains of at least four ubiquitin moieties appear to be required for efficient recognition by the 26S proteasome, although the conformations of the polyubiquitin chains recognized by the proteasome or by other enzymes involved in ubiquitin metabolism are currently unknown. A new crystal form of tetraubiquitin, which has two possible chain connectivities that are indistinguishable in the crystal, is reported. In one possible connectivity, the tetraubiquitin chain is extended and packs closely against the antiparallel neighbor chain in the crystal to conceal a hydrophobic surface implicated in 26S proteasome recognition. In the second possibility, the tetraubiqutitin forms a closed compact structure, in which that same hydrophobic surface is buried. Both of these conformations are quite unlike the structure of tetraubiquitin that was previously determined in a different crystal form [Cook et al. (1994), J. Mol. Biol. 236, 601--609]. The new structure suggests that polyubiquitin chains may possess a substantially greater degree of conformational flexibility than has previously been appreciated.
- Biochemistry Department, University of Utah, 211 MREB, 50 North Medical Drive, Salt Lake City, UT 84132, USA.
Organizational Affiliation: