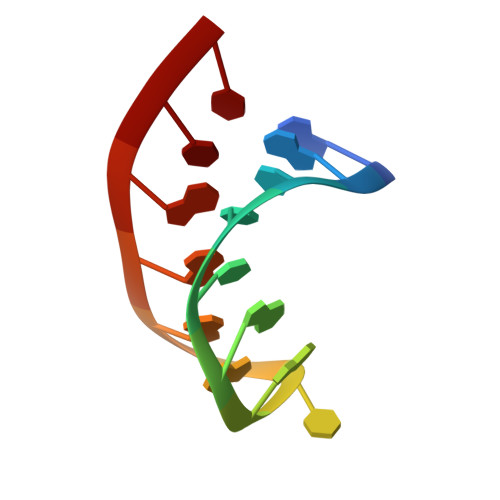
Structures of two RNA domains essential for hepatitis C virus internal ribosome entry site function.
Lukavsky, P.J., Otto, G.A., Lancaster, A.M., Sarnow, P., Puglisi, J.D.(2000) Nat Struct Biol 7: 1105-1110
- PubMed: 11101890 Search on PubMed
- DOI: https://doi.org/10.1038/81951
- Primary Citation Related Structures:
1F84, 1F85 - PubMed Abstract:
Translation of the hepatitis C virus (HCV) polyprotein is initiated at an internal ribosome entry site (IRES) element in the 5' untranslated region of HCV RNA. The HCV IRES element interacts directly with the 40S subunit, and biochemical experiments have implicated RNA elements near the AUG start codon as required for IRES-40S subunit complex formation. The data we present here show that two RNA stem loops, domains IIId and IIIe, are involved in IRES-40S subunit interaction. The structures of the two RNA domains were solved by NMR spectroscopy and reveal structural features that may explain their role in IRES function.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, California 94305-5126, USA.
Organizational Affiliation: