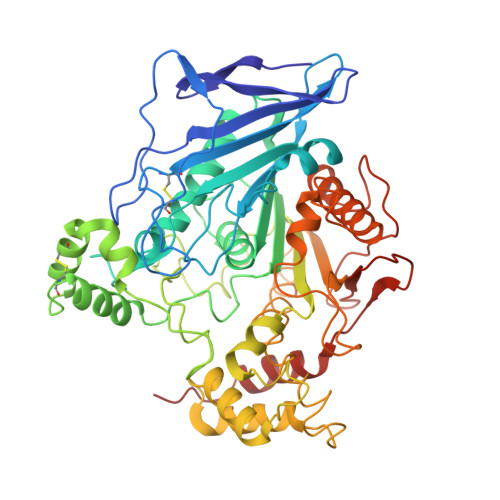
Crystal structure of the catalytic domain of human bile salt activated lipase.
Terzyan, S., Wang, C.S., Downs, D., Hunter, B., Zhang, X.C.(2000) Protein Sci 9: 1783-1790
- PubMed: 11045623 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.9.9.1783
- Primary Citation Related Structures:
1F6W - PubMed Abstract:
Bile-salt activated lipase (BAL) is a pancreatic enzyme that digests a variety of lipids in the small intestine. A distinct property of BAL is its dependency on bile salts in hydrolyzing substrates of long acyl chains or bulky alcoholic motifs. A crystal structure of the catalytic domain of human BAL (residues 1-538) with two surface mutations (N186D and A298D), which were introduced in attempting to facilitate crystallization, has been determined at 2.3 A resolution. The crystal form belongs to space group P2(1)2(1)2(1) with one monomer per asymmetric unit, and the protein shows an alpha/beta hydrolase fold. In the absence of bound bile salt molecules, the protein possesses a preformed catalytic triad and a functional oxyanion hole. Several surface loops around the active site are mobile, including two loops potentially involved in substrate binding (residues 115-125 and 270-285).
- Crystallography Program, Oklahoma Medical Research Foundation, Oklahoma City 73104, USA.
Organizational Affiliation: