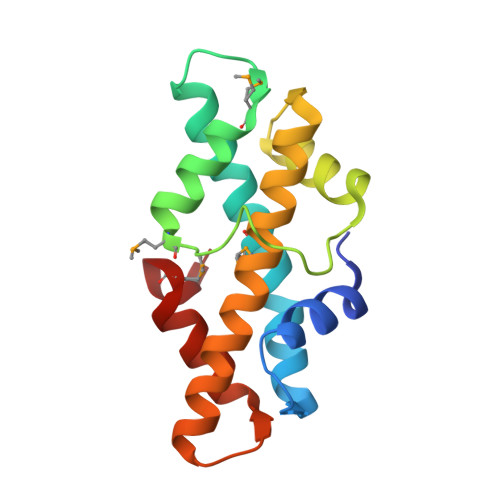
An ancestral nuclear protein assembly: crystal structure of the Methanopyrus kandleri histone.
Fahrner, R.L., Cascio, D., Lake, J.A., Slesarev, A.(2001) Protein Sci 10: 2002-2007
- PubMed: 11567091 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.10901
- Primary Citation Related Structures:
1F1E - PubMed Abstract:
Eukaryotic histone proteins condense DNA into compact structures called nucleosomes. Nucleosomes were viewed as a distinguishing feature of eukaryotes prior to identification of histone orthologs in methanogens. Although evolutionarily distinct from methanogens, the methane-producing hyperthermophile Methanopyrus kandleri produces a novel, 154-residue histone (HMk). Amino acid sequence comparisons show that HMk differs from both methanogenic and eukaryotic histones, in that it contains two histone-fold ms within a single chain. The two HMk histone-fold ms, N and C terminal, are 28% identical in amino acid sequence to each other and approximately 21% identical in amino acid sequence to other histone proteins. Here we present the 1.37-A-resolution crystal structure of HMk and report that the HMk monomer structure is homologous to the eukaryotic histone heterodimers. In the crystal, HMk forms a dimer homologous to [H3-H4](2) in the eukaryotic nucleosome. Based on the spatial similarities to structural ms found in the eukaryotic nucleosome that are important for DNA-binding, we infer that the Methanopyrus histone binds DNA in a manner similar to the eukaryotic histone tetramer [H3-H4](2).
- Department of Chemistry and Biochemistry, University of California, Los Angeles, California 90095-1570, USA.
Organizational Affiliation: