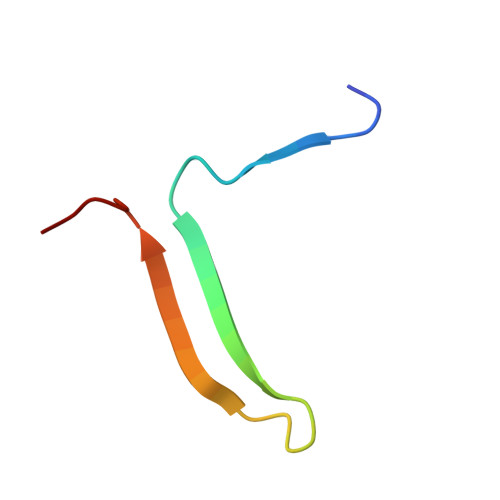
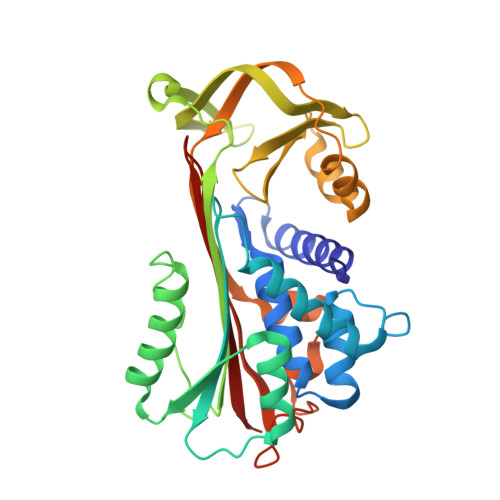
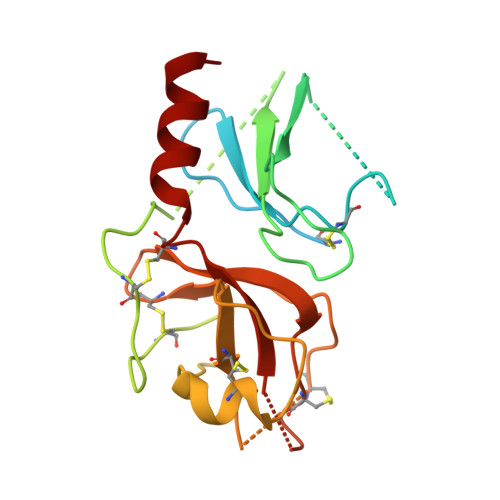
Structure of a serpin-protease complex shows inhibition by deformation.
Huntington, J.A., Read, R.J., Carrell, R.W.(2000) Nature 407: 923-926
- PubMed: 11057674 Search on PubMed
- DOI: https://doi.org/10.1038/35038119
- Primary Citation Related Structures:
1EZX - PubMed Abstract:
The serpins have evolved to be the predominant family of serine-protease inhibitors in man. Their unique mechanism of inhibition involves a profound change in conformation, although the nature and significance of this change has been controversial. Here we report the crystallographic structure of a typical serpin-protease complex and show the mechanism of inhibition. The conformational change is initiated by reaction of the active serine of the protease with the reactive centre of the serpin. This cleaves the reactive centre, which then moves 71 A to the opposite pole of the serpin, taking the tethered protease with it. The tight linkage of the two molecules and resulting overlap of their structures does not affect the hyperstable serpin, but causes a surprising 37% loss of structure in the protease. This is induced by the plucking of the serine from its active site, together with breakage of interactions formed during zymogen activation. The disruption of the catalytic site prevents the release of the protease from the complex, and the structural disorder allows its proteolytic destruction. It is this ability of the conformational mechanism to crush as well as inhibit proteases that provides the serpins with their selective advantage.
- Department of Haematology, University of Cambridge, Wellcome Trust Centre for Molecular Mechanisms in Disease, Cambridge Institute for Medical Research, UK. rwc1000@cam.ac.uk
Organizational Affiliation: