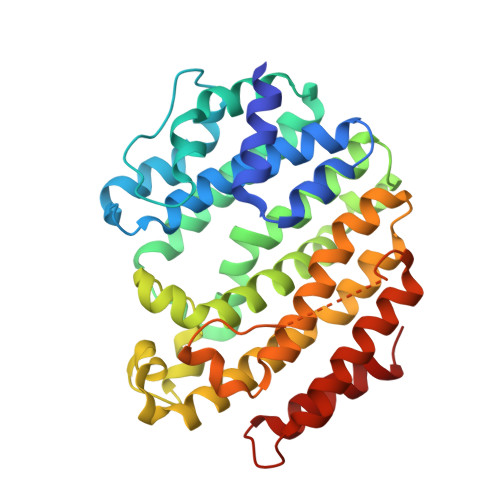
Crystal structure of human squalene synthase. A key enzyme in cholesterol biosynthesis.
Pandit, J., Danley, D.E., Schulte, G.K., Mazzalupo, S., Pauly, T.A., Hayward, C.M., Hamanaka, E.S., Thompson, J.F., Harwood Jr., H.J.(2000) J Biological Chem 275: 30610-30617
- PubMed: 10896663 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M004132200
- Primary Citation Related Structures:
1EZF - PubMed Abstract:
Squalene synthase catalyzes the biosynthesis of squalene, a key cholesterol precursor, through a reductive dimerization of two farnesyl diphosphate (FPP) molecules. The reaction is unique when compared with those of other FPP-utilizing enzymes and proceeds in two distinct steps, both of which involve the formation of carbocationic reaction intermediates. Because FPP is located at the final branch point in the isoprenoid biosynthesis pathway, its conversion to squalene through the action of squalene synthase represents the first committed step in the formation of cholesterol, making it an attractive target for therapeutic intervention. We have determined, for the first time, the crystal structures of recombinant human squalene synthase complexed with several different inhibitors. The structure shows that SQS is folded as a single domain, with a large channel in the middle of one face. The active sites of the two half-reactions catalyzed by the enzyme are located in the central channel, which is lined on both sides by conserved aspartate and arginine residues, which are known from mutagenesis experiments to be involved in FPP binding. One end of this channel is exposed to solvent, whereas the other end leads to a completely enclosed pocket surrounded by conserved hydrophobic residues. These observations, along with mutagenesis data identifying residues that affect substrate binding and activity, suggest that two molecules of FPP bind at one end of the channel, where the active center of the first half-reaction is located, and then the stable reaction intermediate moves into the deep pocket, where it is sequestered from solvent and the second half-reaction occurs. Five alpha helices surrounding the active center are structurally homologous to the active core in the three other isoprenoid biosynthetic enzymes whose crystal structures are known, even though there is no detectable sequence homology.
- Departments of Exploratory Medicinal Sciences and Cardiovascular and Metabolic Diseases, Pfizer Central Research, Groton, Connecticut 06340, USA. jayvardhan_pandit@groton.pfizer.com
Organizational Affiliation: