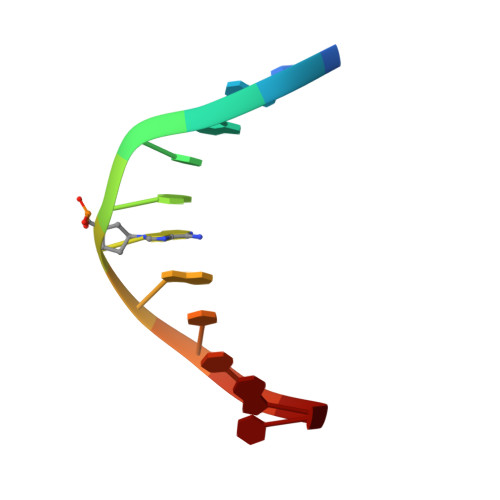
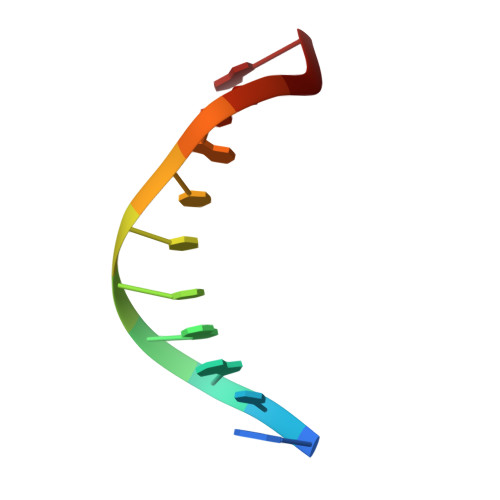
Structure of an 11-mer DNA duplex containing the carbocyclic nucleotide analog: 2'-deoxyaristeromycin
Smirnov, S., Johnson, F., Marumoto, R., de los Santos, C.(2000) J Biomol Struct Dyn 17: 981-991
- PubMed: 10949165 Search on PubMed
- DOI: https://doi.org/10.1080/07391102.2000.10506586
- Primary Citation Related Structures:
1EXL - PubMed Abstract:
2'-deoxyaristeromycin (dAr) is a nucleoside analogue that is resistant to the action of DNA glycosylases. High-resolution NMR spectroscopy and molecular dynamics simulations were used to determine the three-dimensional structure of an 11-mer DNA containing a single dAr.T base pair at its center. Analysis of the spectra revealed the existence of a right-handed duplex in solution, stabilized by Watson-Crick hydrogen bonding and base-stacking interactions. The carbocyclic sugar adopted a C1'-exo conformation and sugars of the 3'-flanking base pair had puckers in the O4'-endo range. The dAr.T base pair was mildly propeller twisted, and the dAr analogue showed a positive roll with the 3'-flanking base. Our findings indicate that the observed resistance of dAr-containing oligodeoxynucleotides to the catalytic action of DNA glycosylases relates to its electronic properties rather than structure, and validate the use of dAr and related carbocyclic nucleoside analogues for biological and structure/function relationship studies.
- Department of Pharmacological Sciences, State University of New York at Stony Brook, 11794-8651, USA.
Organizational Affiliation: