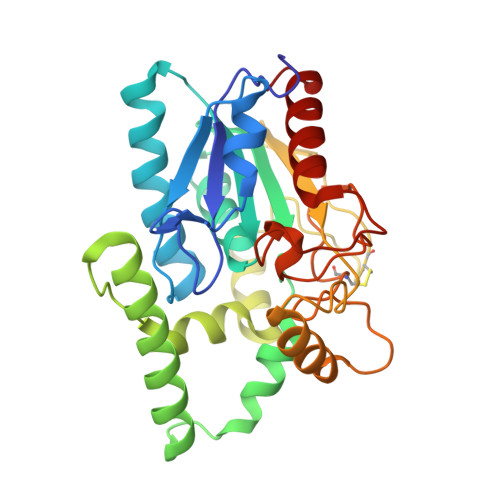
Crystal structure of pseudomonas aeruginosa lipase in the open conformation. The prototype for family I.1 of bacterial lipases.
Nardini, M., Lang, D.A., Liebeton, K., Jaeger, K.E., Dijkstra, B.W.(2000) J Biological Chem 275: 31219-31225
- PubMed: 10893416 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M003903200
- Primary Citation Related Structures:
1EX9 - PubMed Abstract:
The x-ray structure of the lipase from Pseudomonas aeruginosa PAO1 has been determined at 2.54 A resolution. It is the first structure of a member of homology family I.1 of bacterial lipases. The structure shows a variant of the alpha/beta hydrolase fold, with Ser(82), Asp(229), and His(251) as the catalytic triad residues. Compared with the "canonical" alpha/beta hydrolase fold, the first two beta-strands and one alpha-helix (alphaE) are not present. The absence of helix alphaE allows the formation of a stabilizing intramolecular disulfide bridge. The loop containing His(251) is stabilized by an octahedrally coordinated calcium ion. On top of the active site a lid subdomain is in an open conformation, making the catalytic cleft accessible from the solvent region. A triacylglycerol analogue is covalently bound to Ser(82) in the active site, demonstrating the position of the oxyanion hole and of the three pockets that accommodate the sn-1, sn-2, and sn-3 fatty acid chains. The inhibited enzyme can be thought to mimic the structure of the tetrahedral intermediate that occurs during the acylation step of the reaction. Analysis of the binding mode of the inhibitor suggests that the size of the acyl pocket and the size and interactions of the sn-2 binding pocket are the predominant determinants of the regio- and enantio-preference of the enzyme.
- Laboratory of Biophysical Chemistry and BIOSON Research Institute, Department of Chemistry, University of Groningen, Nijenborgh 4, 9747 AG Groningen, The Netherlands.
Organizational Affiliation: